|
|
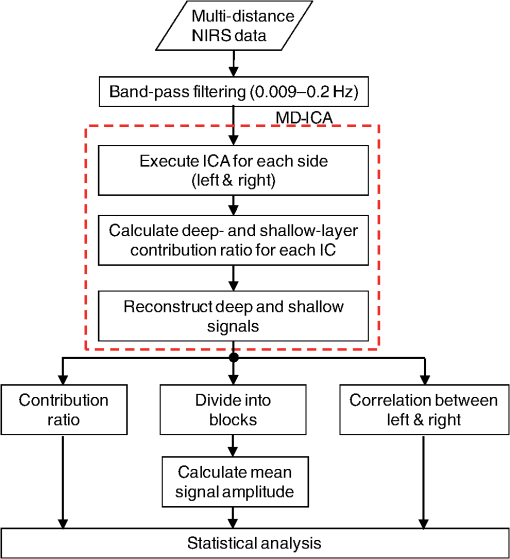
1.IntroductionCerebral blood volume change is measured by radiating weak visible or near-infrared light into the head and detecting the transmitted light from another position on the basis of near-infrared spectroscopy (NIRS).1–5 An NIRS imaging system, such as optical topography, that uses multiple light sources and detectors is used to obtain two-dimensional topographical images of the changes in brain hemodynamics and oxygenation.6–8 NIRS systems have been widely used for research and clinical purposes,9–11 especially for measuring the brain activity of infants,12–17 detecting the functional connectivity on the basis of spontaneous activity of the brain of infants while resting,18–21 and using wearable systems to conduct measurements in everyday situations.22,23 The general consistency between NIRS signals and blood oxygenation level dependent signals has been shown in studies on task-evoked brain activation and resting-state functional connectivity,24–26 validating NIRS as a neuroimaging tool. However, a major issue for NIRS is to what extent the deep tissue (such as cerebral tissue), as opposed to shallow-tissue (extracerebral tissue, such as the scalp), contributes to the measured signals of oxygenated and deoxygenated hemoglobin (oxy- and deoxy-Hb). Task-evoked and/or spontaneous changes in skin blood flow due to systemic effects can mask changes in cerebral oxygenation in relation to neural activation.27–31 Previous studies have examined the effects of the shallow tissue on NIRS signals in adults by means of additional channels with short source-detector (S-D) distances under the assumption that the short distance channels measure signals only from the shallow tissue.32,33 Time-domain NIRS has also been used to separate deep- from shallow-tissue signals.34,35 Simultaneous recordings of NIRS with cutaneous laser-Doppler blood flow have shown that the scalp blood flow contributes to the changes in NIRS signals.29,36 The separation of deep- from shallow-tissue signals has also been performed by time-series analysis, such as adaptive filtering37 or independent component analysis (ICA).38 Most of the aforementioned studies assumed either the spatially homogeneous properties of systemic effects on the shallow tissues or the independence of temporal evolution of the deep and shallow signals. However, the assumptions might be violated if the systemic effects induced by tasks can locally affect the blood flow and oxygenation in both deep- and shallow tissues. Kirilina et al.39 used NIRS, functional magnetic resonance imaging (fMRI), MR-angiography, blood pressure, heart rate, skin conductance, and skin blood flow to study activation in the frontal cortex during a cognitive task and demonstrated that cognitive tasks strongly affect the blood volume of veins draining the scalp and that NIRS signals contain the task-evoked systemic artifacts. Funane et al.40 proposed a method to discriminate between the deep- and shallow-tissue effects on the NIRS signal using multiple-distant (MD) probes and an ICA.41,42 Importantly, the MD-ICA method can be used to decompose NIRS signals even when the cerebral and extracerebral signals correlate. While NIRS studies have provided unique opportunities to reveal the developmental processes of the functional activation in the brains of infants12–17 and functional connectivity,19–21 very few have examined how deep- (e.g., cerebral) and shallow-tissue (scalp) layers of the head contribute to NIRS signals. Since the hemodynamic responses to visual or auditory stimuli in young infants show global properties over diverse cortical regions,43,44 it is crucial to elucidate the systemic effects on the hemodynamic responses for both shallow and deep tissues. Moreover, the dissociation of deep and shallow signals is important in the study of the functional connectivity of the cortex in the resting state, because a false-positive connectivity might be estimated if measured NIRS signals have a high contribution from the shallow tissue that exhibits temporally correlated hemodynamic changes over different locations of the scalp. It is also important to note that infants’ heads differ from those of adults not only in size, but also for optical properties regarding NIR light.45 Thus, an extrapolation from adult studies may not be valid. Taga et al.46 investigated the effect of the source-detector (S-D) distance on signal amplitude using MD probes with infants, but no method has been applied for separating shallow and deep signals. The MD-ICA method40 can be applied to the infant data obtained using MD probes if infant-specific parameters are determined, such as optical path length in each head tissue45,47 and brain-scalp distance.48 This study is intended to investigate the cerebral and extracerebral contributions to NIRS signals during functional activation in infants. Furthermore, we also investigate those during a resting state, which has not been shown in the previous study.40 To do this, we develop a model to determine infant-specific parameters for optical properties and use the MD-ICA method,40 which was verified for adult participants, to quantify the effect of deep and shallow tissue on NIRS signals measured with infants’ heads in the existing data.46 2.Method2.1.ParticipantsThe data used in this study were originally obtained for a previous study46 in which nine three-month-old (mean age, 113 days) infants (six girls and three boys) participated. All infants were studied while they were quietly sleeping. Informed consent was obtained from the parents of the infants prior to the initiation of the experiments. All experiments were approved by the Office for Life Science Research Ethics and Safety, The University of Tokyo. 2.2.NIRS MeasurementAn NIRS system (ETG-100, Hitachi Medical Corporation, Tokyo, Japan)6,7 was used to measure the hemodynamic responses of infants. The optical fibers for the source and detectors were placed over the temporal cortices on both hemispheres. On the left hemisphere, the source was set on the T3 position on the basis of the international 10–20 system of electrode placement,49 and four detectors were mounted in a line at 10, 20, 30, and 40 mm from the source. On the right hemisphere, the source and four detectors were placed on homologous positions. Figure 1 shows the arrangement of the source and detectors in NIRS measurements for hemodynamic responses to auditory stimuli in infants. Thus all the measurement channels in the present study were likely located on the temporal cortex.16,50 These regions have also been reported to respond to speech sounds while infants are quietly sleeping.51 Fig. 1Arrangement of the light sources and detectors in near-infrared spectroscopy (NIRS) measurements for hemodynamic responses to auditory stimuli in infants. Filled and open circles show the positions of the light sources and detectors, respectively. The source optical fibers were placed on the T3 and T4 positions of the International 10/20 system. The position of the measurement channel for recording the changes in oxygenated and deoxygenated hemoglobin (oxy- and deoxy-Hb) signals is defined as the midpoint between the source and detector. Four different detectors placed in a line on each hemisphere detect reflected light from a single incident position. Thus, hemoglobin signals are simultaneously measured by a total of eight channels. The data obtained at source-detector 40-mm channel were not used in the analysis. 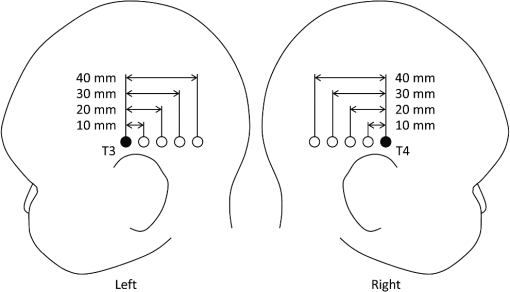 Although in the previously obtained data46 two different intensities of NIR light were used, i.e., 0.6 mW (0.3 mW for each of the two laser diodes; 780 and 830 nm) and 1.2 mW (0.6 mW for each), in this study only the dataset measured under 1.2 mW was used because the signal-to-noise ratio of the detected light intensity is higher. The data obtained at 40-mm S-D distance were not used for the analysis because signal-to-noise ratios for several participants were not sufficient for applying the MD-ICA method. 2.3.StimulationDetails of the stimulation have been described in previous papers.46,52 The test auditory stimuli consisted of speech sounds of Japanese words provided by a female Japanese speaker. Ten different sets of stimuli (mean duration: 2.68 s; range: 2.54 to 2.81 s) were used. The experiments were conducted in a sound-attenuated room. The infants sat on the lap of an experimenter and were studied while they were quietly sleeping. The speech sounds were presented at a 73-dB maximum amplitude. We used an event-related paradigm under the speech condition in which the test auditory stimuli were presented every 15 s (i.e., stimulus onset ). During interstimulus intervals, we did not present any sound. The measurements lasted 150 s. Under the no-stimulus (control) condition, the infants were given no sound and were measured for 150 s. 2.4.Preprocess for NIRS DataFor the preprocessing for the NIRS data analysis, the oxy- and deoxy-Hb changes were calculated by using the optical density changes of 780- and 830-nm light in accordance with the modified Beer-Lambert law.6,53 The analysis flow of this study is shown in Fig. 2. We applied a 0.009 to a 0.2 Hz band-pass filter for suppressing the pulse signals and low-frequency fluctuation. The blocks were not used where the nonfiltered (raw) oxy-Hb signal changed very quickly (over change in 1 s) at either the S-D 10- or 20-mm channel because such blocks probably contain some artifacts. For the MD-ICA method, continuous data, including blocks, are necessary for applying ICA. If a block was eliminated, the blocks after or before the eliminated block all had to be eliminated, leaving longer continuous data that did not include the eliminated blocks. With this procedure, out of 180 blocks (), eight were not used. In individual datasets, we initially divided the continuous time course data into blocks. Each data block ranged from 1 s prior to stimulus onset to 15 s after onset. Then, we corrected the baseline using linear fitting to the mean signal of the first 1 s and that of the last 1 s in each data block. By averaging the signal changes over the data blocks for each subject under each condition, we obtained individual hemodynamic responses for each measurement channel. We finally obtained group-averaged hemodynamic responses at each measurement channel under each condition. 2.5.Deep-Shallow Signal Separation MethodThere are several methods for removing the scalp effect from an NIRS signal.40,54,55 We previously proposed a method with MD probes to discriminate between the deep- (such as cerebral) and shallow- (scalp) tissue effects on the NIRS signals obtained from adult participants.40 We base the present study on the previous work and apply the previously proposed method to infant data. In the method with MD-ICA, it is assumed that the partial optical path length of the deep layer linearly increases as the S-D distance from increases, whereas that of the shallow layer does not change. indicates the shortest S-D distance at which the detected light is sensitive to absorption change in gray matter. This is assumed to be 10.5 mm in adults (experimentally obtained).40 For infants, a different value, i.e., 8 mm, was used as mentioned below. Moreover, we assumed that the NIRS signals at each S-D distance can be expressed by the linear sum of the hemoglobin-change signals that are proportional to the partial optical path length at scalp and gray matter.56 In the first step of this method, an ICA41,42 is used for obtaining signal sources (independent components) to acquire the deep and shallow subcomponents described in the following step. In this step of the analysis, three independent components were obtained for each hemisphere. If enough (e.g., four or more) channels are available, noise components can generally be truncated. However, we did not cut any components in the MD-ICA analysis because not enough channels were available. In the next step, the contribution ratios of the deep and shallow layers are calculated by the dependence of signal amplitudes (i.e., weight of component) on the S-D distance for each independent component. The deep and shallow subcomponents are then calculated by multiplying the independent components by deep/shallow contribution ratios. In the final step, deep and shallow signals are reconstructed using the linear sum of the subcomponents of all the independent components. This analysis supports a model that allows us to calculate an estimation of the relative contributions. To explain in more detail, the deep and shallow signals are reconstructed by the following equations: where to Pm indicate position numbers, to Cn indicate component numbers, indicates the hemoglobin change (product of concentration and optical path length), and indicate deep (dp) and shallow (sh) signals, indicates the time course of the independent component (Cn), and indicate deep and shallow layer contribution ratios for component Cn, and indicates the weight of component Cn as the output of the ICA at position Pm. When all independent components are used, can be expressed asTo reconstruct deep and shallow signals, weight () and contribution ratio () are used. is obtained with the ICA process and is obtained with the -intercept using a product of weight and the effective amplitude of the component along with multiple S-D distances. The effective amplitude of the component is calculated using the root mean square of the time series of the component. Figure 3 shows a geometric explanation for calculating the contribution ratio () of deep and shallow signals in the original NIRS signal (total signal including deep- and shallow-tissue contributions) under two -intercept conditions. A line, obtained by at least two signal contributions (product of and the effective component amplitude) is used to obtain an -intercept (). indicates the experimentally obtained -intercept for an independent component that includes both deep- and shallow-layer effects. A pure deep (brain) component should have as an -intercept, and the shallow-layer contribution is constant for all S-D distance channels. By using this model, the deep-layer contribution ratio can be calculated by when is shorter than [Fig. 3(a)] and by when is longer than [Fig. 3(b)]. When a component signal is included in both deep and shallow signals, would be shorter than [Fig. 3(a)] because contributions from both layers are in the same direction as the sign. Adversely, when a component is positively included in the deep signal and negatively included in the shallow signal (or vice versa), would be longer than [Fig. 3(b)]. Fig. 3Geometric explanation for calculation of contribution ratios () of deep and shallow signals in original NIRS signal under two -intercept conditions: (a) and (b) . The deep-layer contribution ratios for component Ci are expressed as in (a) and in (b). 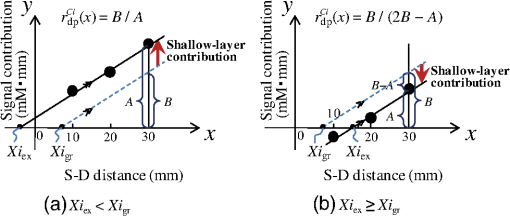 To confirm the assumption of a partial optical path length for deep and shallow layers in an infant’s head, we performed a Monte Carlo simulation for photon propagation in an infant’s head under conditions of 10, 20, and 30-mm S-D distances. Absorption and reduced scattering coefficients of the neonatal head model were adopted from Fukui et al. (Table 1 in Ref. 45). The thickness of each tissue layer was proportionally converted from the data of Okada and Delpy (Table 1 in Ref. 57) so that the scalp surface and gray matter surface were 8.5 mm apart (Fig. 3A in Ref. 48). Figure 4 shows partial optical path lengths for scalp and gray matter layers for infants calculated from the Monte Carlo simulation (number of detected photons: 10,000). We assumed that the difference between the of neonates and three-month-old infants is small enough (considering the difference between the thickness of each layer for neonates and three-month-old infants as reported by Beauchamp et al.).48 Table 1Deep-layer contribution ratio.
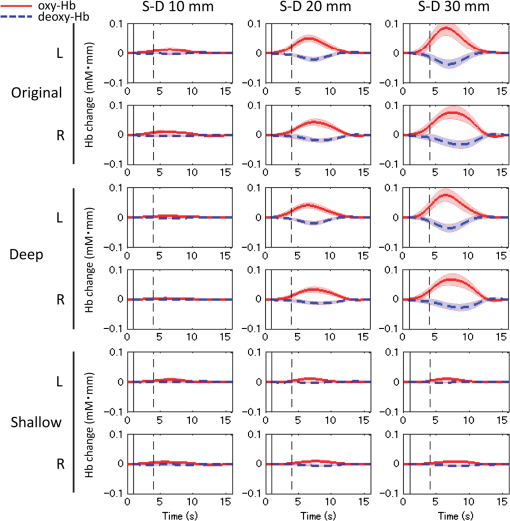
S-D, source-detector; oxy-Hb, oxygenated hemoglobin; deoxy-Hb, deoxygenated hemoglobin. Fig. 4Partial optical path lengths for scalp and gray matter in infant head model by Monte Carlo simulation. 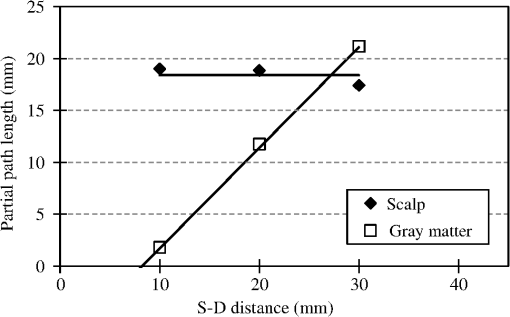 From the simulation, the optical path length in the scalp does not change much between 10- and 30-mm S-D distances, while the optical path length for the gray matter linearly increases. The results were similar to those of simulations performed on an adult head model.40,58,59 Therefore, the necessary assumption of MD-ICA was validated. The results also showed that the is 8 mm for the infant head model. In the following, data are analyzed using 8 mm as the . The shortest S-D distance used in this study is 10 mm, which is longer than . Even if an S-D distance shorter than is used (e.g., 5 mm) and a pure scalp signal is obtained, the contributions of shallow (scalp) and deep (cerebral) signals cannot be correctly quantified when the signals are similar (temporally correlated with each other). To overcome the limitation, we used the dependence of signal components on the S-D distance using multidistance optodes and ICA, where multiple S-D distances longer than the are used. 2.6.Analysis After Deep-Shallow SeparationThe following analyses were conducted after deep and shallow signals were reconstructed from the original NIRS signals (Fig. 2). 2.6.1.Deep-layer contribution ratioMeans and standard deviations of deep-layer contribution ratios for each S-D distance of left and right channels were calculated by the MD-ICA method. We performed a three-way analysis of variance (ANOVA) [S-D distance (no stimulus/speech)] for the deep-layer contribution ratios of oxy- and deoxy-Hb signals. The statistical threshold was set at in order to take into account multiple comparisons between the two types of Hb signals. 2.6.2.Mean signal amplitude of blocksWe divided each continuous dataset into block data that included 1 s before the onset time to 12 s after the stimulation for each trial. After we divided the continuous data into blocks, the mean amplitude during a time window (4.1 to 7.0 s from onset for oxy-Hb and 5.0 to 7.6 s from onset for deoxy-Hb) was calculated. The time window was adopted from that of Taga et al.46 We performed a three-way ANOVA [S-D distance (original/deep/shallow)] for the mean signal amplitude under the no-stimulus and speech conditions. The statistical threshold was set at in order to take into account multiple comparisons between the two types of Hb signals under the two conditions. 2.6.3.Correlation coefficients between left and right channelsThe functional connectivity among the cortical regions has been assessed by calculating the temporal correlations of the continuous oxy- and deoxy-Hb signals obtained at multichannels under no-stimulus conditions.19,21 To investigate the interhemispheric functional connectivity between the homologous regions of the temporal cortices and the effect of the superficial layer on the connectivity, the correlation coefficients between the left-right channels at each S-D distance were calculated for original, deep, and shallow signals. By using continuous data from original, deep, and shallow signals, correlation coefficients (Fisher’s )60,61 between the left-right channels were calculated at each S-D distance and task condition. The results were compared among all conditions. We performed a three-way ANOVA [S-D distance (no stimulus/speech)] for the correlation coefficients of oxy- and deoxy-Hb signals. The statistical threshold was set at in order to take into account multiple comparisons between the two types of Hb signals. 3.Results3.1.Block-Averaged NIRS Time CoursesBlock-averaged NIRS time courses for each layer (original/deep/shallow) under the speech condition are shown in Fig. 5. Red solid and blue dashed lines indicate oxy- and deoxy-Hb, respectively. Vertical solid and dashed lines indicate the start (onset) and end of the stimulation period, respectively. 3.2.Deep-Layer ContributionTable 1 lists means and standard deviations among participants of deep-layer contribution ratios for the left and right channels under the no-stimulus and speech conditions. Table 2 lists three-way ANOVA results for deep-layer contribution ratio ( values). S-D distance had a significant main effect on deep-layer contribution ratios of both oxy- [, ] and deoxy-Hb [, ] signals. A post hoc test (Tukey-Kramer) was conducted to identify specific differences among the contribution ratios obtained at the three S-D distances. In the results for both oxy- and deoxy-Hb signals, the mean of the contribution ratios at each S-D distance was significantly different from those at the other S-D distances (). Table 2Three-way analysis of variance (ANOVA) results (F values) for deep-layer contribution ratio.
The numbers in parentheses in the first column indicate the degrees of freedom for each factor and interaction. Single (*), double (**), and triple (***) asterisks denote statistical significance at p<0.025, 0.005, and 0.0005, respectively. The test conditions (no stimulus/speech) had significant main effects on the deep-layer contribution ratios of both oxy-Hb [, ] and deoxy-Hb [, ] signals. Interactions between hemispheres and conditions were significant for both oxy-Hb () and deoxy-Hb () signals. 3.3.Mean Signal AmplitudeMean signal amplitudes for each condition are shown in Fig. 6. In the data obtained at S-D 20 and 30 mm, original- and deep-layer signals showed similar changes in mean signal amplitudes. In the data obtained at S-D 10 mm, which included relatively higher scalp effects than other longer S-D distances, more significant signal changes in oxy- and deoxy-Hb were obtained in the deep signal than in the original and shallow signals. No significant change in the mean signal amplitude was obtained under the no-stimulus condition. Fig. 6Mean signal amplitude. (a) Original oxy-Hb, (b) original deoxy-Hb, (c) deep oxy-Hb, (d) deep deoxy-Hb, (e) shallow oxy-Hb, and (f) shallow deoxy-Hb signals. Single (∗), double (∗∗), and triple (∗∗∗) asterisks denote statistical significance at , 0.01, and 0.005, respectively. 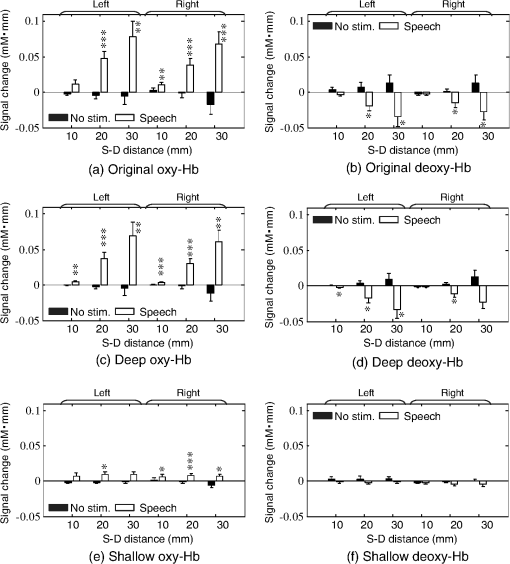 Three-way ANOVA results ( values) for the mean signal change are shown in Table 3. Under the speech condition, the S-D distance had a significant main effect on the mean signal change of both oxy-Hb [, ] and deoxy-Hb [, ] signals. A post hoc test (Tukey-Kramer) was conducted to identify specific differences among mean signal changes obtained at the three S-D distances. In the results, the mean signal change at each S-D distance was significantly different from those at the other two S-D distances () only for the oxy-Hb signals. The layer value (original/deep/shallow) also had a significant main effect on the mean signal change of both oxy-Hb [, ] and deoxy-Hb [, ] signals. The results of a post hoc test (Tukey-Kramer) show that the absolute value of the shallow signal was significantly smaller than those of the other (original and deep) signals ( for oxy-Hb, for deoxy-Hb). Interactions between the S-D distance and layer (original/deep/shallow) were significant only for the oxy-Hb signal (). Table 3Three-way ANOVA results (F values) for mean signal change.
The numbers in parentheses in the first column indicate the degrees of freedom for each factor and interaction. Triple (***) asterisks denote statistical significance at p<0.0005. Under the no-stimulus condition, no factor had a significant main effect on the mean signal change of either oxy- or deoxy-Hb (). Interactions between the three factors were not significant (). 3.4.Correlation Between Left-Right ChannelsCorrelation coefficients (Fisher’s ) between the left and right channels for all participants are shown in Fig. 7. Bar graphs indicate means among the participants. Table 4 lists the three-way ANOVA results ( values) for correlation coefficients between the left and right channels. The S-D distance had a significant main effect on correlation coefficients for both oxy-Hb [, ] and deoxy-Hb [, ] signals. For both cases of oxy- and deoxy-Hb signals, post hoc tests (Tukey-Kramer) showed that correlation coefficients of the S-D 10-mm data were significantly smaller than those of the other S-D distance data (20 and 30 mm) ( for both oxy- and deoxy-Hb), whereas there was no difference between the correlation coefficients of the S-D 20- and 30-mm data ( for both oxy- and deoxy-Hb). Fig. 7Correlation coefficients () between left and right channels for all participants. Bar graphs indicate means among participants. 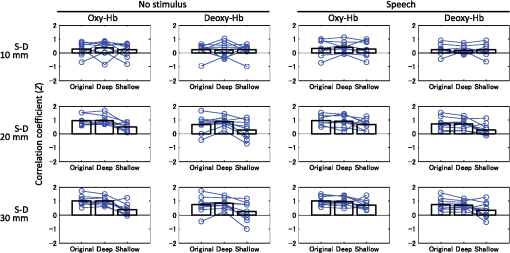 Table 4Three-way ANOVA results (F values) for correlation coefficients between left and right channels.
The numbers in parentheses in the first column indicate the degrees of freedom for each factor and interaction. Double (**) and triple (***) asterisks denote statistical significance at p<0.005 and 0.0005, respectively. The type of layer (original/deep/shallow) had a significant main effect on correlation coefficients for both oxy-Hb [, ] and deoxy-Hb [, ] signals. For both cases of oxy- and deoxy-Hb signals, post hoc test results showed that correlation coefficients of the shallow signals were significantly smaller than those of the other layer (original and deep) signals ( for oxy-Hb, for deoxy-Hb), whereas there was no difference between the correlation coefficients of original and deep signals ( for both oxy- and deoxy-Hb). 4.Discussion4.1.Deep-Layer Contribution RatioThe present study analyzing data of three-month-old infants showed that the deep-layer contribution ratios (averaged between left and right channels) at a 30-mm S-D distance under the speech condition for oxy- and deoxy-Hb were 82 and 78%, respectively, as shown in Table 1. Note that Funane et al.40 reported that in adults, the mean deep-layer contribution ratios at the S-D 30-mm channels for oxy- and deoxy-Hb were 65 and 67% among several tasks, respectively. Thus, the deep-layer effect in infants was greater than that in adults. For 20-mm channels, deep-layer contribution ratios (averaged between left and right channels) under the speech condition for oxy- and deoxy-Hb were 72 and 66%, respectively, while for the 10-mm channels, deep-layer contribution ratios were . Thus, a significant amount of signal originates from deep (such as cerebral) tissue at 20- and 30-mm S-D distances. This was also the case under the no-stimulus condition, suggesting that NIRS signals largely reflect spontaneous changes in cerebral blood oxygenation. Because the deep-layer contribution ratio is high enough for both 20- and 30-mm channels, the optimal S-D distance depends on a trade-off between probe density and the area covered by a limited number of optodes. If the probe density needs to be high, the S-D distance should be 20 mm, whereas if the covered probe area needs to be larger, the S-D distance should be 30 mm, for example. According to Table 2, the interaction between hemisphere and condition was significant for both oxy- and deoxy-Hb. This was possibly because the contribution of the deep signal induced by stimulation may be larger in the channel for a large response than that in the channel for a smaller response in an infant’s head. The deep-layer contribution of the oxy-Hb signal is slightly more than that of the deoxy-Hb signal. Franceschini et al.,62 on the other hand, reported that oxy-Hb may be more altered by systemic changes than the deoxy-Hb in adult participants. If the systemic signal contribution is significant, the deep-layer contribution should decrease because such systemic signals are included in both cerebral and extracerebral tissues as reported by Scholkmann et al.9 Therefore, in infants, the extracerebral effect might be smaller than that in adults. Nonetheless, the task dependency of extracranial and systemic effects on NIRS signals in infants is an important issue30 to be further investigated. 4.2.Effect of Deep-Shallow Separation on Mean Signal AmplitudeAs shown in Fig. 6, at S-D 20 and 30 mm, original and deep layer signals in oxy- and deoxy-Hb showed similar changes in mean signal amplitudes. For S-D 10-mm channel, where the most scalp effects are included among the three S-D distances, more significant signal changes in oxy-Hb were obtained in deep signals than in original signals. Significant signal changes were obtained only for the deep signal, especially for the left channel. From this result, MD-ICA effectively extracts a deep signal from infants’ NIRS signals when it is assumed that the deep signal (i.e., such as a cerebral signal) is induced during the stimulation. The amplitude of deep and shallow signals has the dependence on S-D distance that is expected from optical path length. For example, the shallow signal amplitude does not change much as the S-D distance is changed [Figs. 6(e) and 6(f)]. This result demonstrated that the assumed model for infants was valid enough to extract deep and shallow signals. In the ANOVA results, the significant interactions between the S-D distance and type of layer (original/deep/shallow) existed for the oxy-Hb signal because the signal obtained at a longer S-D distance includes a larger contribution from the deep layer based on optical partial path length. Under the speech condition, the deoxy-Hb signal at the S-D 10-mm channel did not show any significant change (only the left deep signal [Fig. 6(d)] is significant). This was not the case for the oxy-Hb signal. This result can be partly explained as a result of the signal amplitude of the deoxy-Hb signal being smaller than that of the oxy-Hb when comparing their variances, as shown in Fig. 5 (original, S-D 10-mm data). For both oxy- and deoxy-Hb signals, on the other hand, the significance of the changes was increased from original to deep signals ( values were decreased). This means that in deep signals, even though the signal amplitudes were lower than those of the original signals, the significance of the signal change increased, which was caused by a reduced signal variance. It has also been shown that the variance in the S-D 10-mm original signal during the set time window is largely caused by the shallow signal. There are some differences between Figs. 6(e) and 6(f). Specifically, significant signal changes were obtained only for the oxy-Hb signal under the speech condition. According to Table 1, however, the deep-layer contribution of the oxy-Hb signal is higher than that of the deoxy-Hb signal. The signal amplitudes were calculated using only a task period (block data), whereas the deep-layer contribution reflects all measurement periods (continuous data). That result, therefore, suggests that low-frequency physiological fluctuations other than a task-related signal may have more effect on deep signals for the deoxy-Hb signal than those for the oxy-Hb signal. In other words, the task-related signal contributes more to the oxy-Hb signal than the deoxy-Hb signal. On the other hand, for deep signals [Figs. 6(c) and 6(d)], larger amplitudes and higher significant changes were obtained for oxy-Hb than for deoxy-Hb, which explains the higher deep-layer contribution for the oxy-Hb signal from another standpoint. In principle, the shallow signal reconstructed by the MD-ICA method mainly reflects light absorption changes in the scalp, and the deep signal reflects those in tissue other than the scalp. From the results of the mean amplitude, the stimulation-related response cannot be explained only using the shallow signal. Therefore, the deep signal mainly originates from cerebral tissue when the blood change in the extracerebral layers in a relatively deeper area (e.g., blood vessels on the brain surface) is negligible. Note that cerebral (originated from local neuronal activity) and systemic signals are not separated by the deep-shallow signal separation method (MD-ICA) because the method only uses signal depth on the basis of optical path length. 4.3.Interhemispheric Functional ConnectivityANOVA and post hoc test results of layer (original/deep/shallow) effects showed that correlation coefficients of the original signals mainly reflect the correlation coefficients of the deep signals and that the left-right correlation cannot be accounted for only by the shallow layer. The significance of the S-D distance main effects also suggests that deep-layer signals largely contribute to the correlation coefficient between the left and right channels because the left-right correlation coefficients are significantly smaller at the S-D 10-mm channel than at other S-D distances. These results ensure that the functional connectivity obtained in infants in the previous study19 largely reflects correlations of deep (such as cerebral) hemodynamic changes. 4.4.LimitationThe infant head model was adopted from the literature45,57 and our original simulation, and was assumed to be 8 mm for all participants. The effects of the model error and linearity of the amplitude of the gray matter signal still need to be investigated, as it does in the case for the application of the MD-ICA method to adults. In this study, one light source is shared by four detectors at one side; therefore, the measurement channel of each detector does not reflect the same position. That is, the channel of the further detector reflects changes observed in the closer detectors and those in the more posterior region in the head. In the MD-ICA method, the homogeneous absorption change in each layer (deep or shallow) is assumed, so the displacement of the channel positions may affect the analytical result and should be as small as possible. Since this study focused only on the temporal cortex of the brain, the shallow-layer effect at other cortices of the brain should be further investigated. The shortest S-D distance at which the detected light is sensitive to absorption change in gray matter depends on the extracerebral tissue thickness, which has a variance around the head. It is, thus, necessary in the future to use structural data (extracerebral tissue thickness) for target areas. 5.ConclusionThe MD-ICA method was applied to dissociate deep- and shallow-layer contributions to NIRS signals obtained in infants and to investigate the left-right connectivity for original, deep, and shallow signals under no-stimulus (resting) and speech conditions. The results revealed that deep tissues contributed more than shallow ones to the NIRS signals when 20- and 30-mm S-D distances were chosen. The deep layers of the NIRS signals in infant heads contributed more than those in adult heads, the results of which were previously reported. Moreover, the original signal correlation between the left-right channels mainly originates from deep signal correlation. These results suggest that the cerebral contribution to NIRS signals is significant in functional activation and connectivity studies of infants. AcknowledgmentsWe thank Kayo Asakawa for her support in the data acquisition and Yoshiko Koda and Keiko Hirano for their administrative assistance. This work was partly supported by Grants-in-Aid for Scientific Research to G.T. (Nos. 20670001, 24119002, and 26220004). ReferencesF. F. Jöbsis,
“Noninvasive, infrared monitoring of cerebral and myocardial oxygen sufficiency and circulatory parameters,”
Science, 198
(4323), 1264
–1267
(1977). http://dx.doi.org/10.1126/science.929199 SCIEAS 0036-8075 Google Scholar
B. Chanceet al.,
“Cognition-activated low-frequency modulation of light absorption in human brain,”
Proc. Natl. Acad. Sci. U. S. A., 90
(8), 3770
–3774
(1993). http://dx.doi.org/10.1073/pnas.90.8.3770 1091-6490 Google Scholar
Y. HoshiM. Tamura,
“Detection of dynamic changes in cerebral oxygenation coupled to neuronal function during mental work in man,”
Neurosci. Lett., 150
(1), 5
–8
(1993). http://dx.doi.org/10.1016/0304-3940(93)90094-2 NELED5 0304-3940 Google Scholar
T. Katoet al.,
“Human visual cortical function during photic stimulation monitoring by means of near-infrared spectroscopy,”
J. Cereb. Blood Flow Metab., 13
(3), 516
–520
(1993). http://dx.doi.org/10.1038/jcbfm.1993.66 JCBMDN 0271-678X Google Scholar
A. Villringeret al.,
“Near infrared spectroscopy (NIRS): a new tool to study hemodynamic changes during activation of brain function in human adults,”
Neurosci. Lett., 154
(1–2), 101
–104
(1993). http://dx.doi.org/10.1016/0304-3940(93)90181-J NELED5 0304-3940 Google Scholar
A. Makiet al.,
“Spatial and temporal analysis of human motor activity using noninvasive NIR topography,”
Med. Phys., 22
(12), 1997
–2005
(1995). http://dx.doi.org/10.1118/1.597496 MPHYA6 0094-2405 Google Scholar
Y. Yamashitaet al.,
“Near-infrared topographic measurement system: imaging of absorbers localized in a scattering medium,”
Rev. Sci. Instrum., 67
(3), 730
–732
(1996). http://dx.doi.org/10.1063/1.1146849 RSINAK 0034-6748 Google Scholar
H. Koizumiet al.,
“Higher-order brain function analysis by trans-cranial dynamic near-infrared spectroscopy imaging,”
J. Biomed. Opt., 4
(4), 403
–413
(1999). http://dx.doi.org/10.1117/1.429959 JBOPFO 1083-3668 Google Scholar
F. Scholkmannet al.,
“A review on continuous wave functional near-infrared spectroscopy and imaging instrumentation and methodology,”
NeuroImage, 85
(1), 6
–7
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.05.004 NEIMEF 1053-8119 Google Scholar
M. FerrariV. Quaresima,
“A brief review on the history of human functional near-infrared spectroscopy (fNIRS) development and fields of application,”
NeuroImage, 63
(2), 921
–935
(2012). http://dx.doi.org/10.1016/j.neuroimage.2012.03.049 NEIMEF 1053-8119 Google Scholar
R. Takizawaet al.,
“Neuroimaging-aided differential diagnosis of the depressive state,”
NeuroImage, 85
(1), 498
–507
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.05.126 NEIMEF 1053-8119 Google Scholar
M. Peñaet al.,
“Sounds and silence: an optical topography study of language recognition at birth,”
Proc. Natl. Acad. Sci. U. S. A., 100
(20), 11702
–11705
(2003). http://dx.doi.org/10.1073/pnas.1934290100 1091-6490 Google Scholar
G. Tagaet al.,
“Brain imaging in awake infants by near-infrared optical topography,”
Proc. Natl. Acad. Sci. U. S. A., 100
(19), 10722
–10727
(2003). http://dx.doi.org/10.1073/pnas.1932552100 1091-6490 Google Scholar
T. Nakanoet al.,
“Prefrontal cortical involvement in young infants’ analysis of novelty,”
Cereb. Cortex, 19
(2), 455
–463
(2009). http://dx.doi.org/10.1093/cercor/bhn096 53OPAV 1047-3211 Google Scholar
J. Gervainet al.,
“Near-infrared spectroscopy: a report from the McDonnell infant methodology consortium,”
Dev. Cogn. Neurosci., 1
(1), 22
–46
(2011). http://dx.doi.org/10.1016/j.dcn.2010.07.004 1878-9293 Google Scholar
H. Watanabeet al.,
“Effect of auditory input on activations in infant diverse cortical regions during audiovisual processing,”
Hum. Brain Mapp., 34
(3), 543
–565
(2013). http://dx.doi.org/10.1002/hbm.21453 HBRME7 1065-9471 Google Scholar
F. Homae,
“A brain of two halves: insights into interhemispheric organization provided by near-infrared spectroscopy,”
NeuroImage, 85
(1), 354
–362
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.06.023 NEIMEF 1053-8119 Google Scholar
G. Tagaet al.,
“Spontaneous oscillation of oxy- and deoxy- hemoglobin changes with a phase difference throughout the occipital cortex of newborn infants observed using non-invasive optical topography,”
Neurosci. Lett., 282
(1–2), 101
–104
(2000). http://dx.doi.org/10.1016/S0304-3940(00)00874-0 NELED5 0304-3940 Google Scholar
F. Homaeet al.,
“Development of global cortical networks in early infancy,”
J. Neurosci., 30
(14), 4877
–4882
(2010). http://dx.doi.org/10.1523/JNEUROSCI.5618-09.2010 JNRSDS 0270-6474 Google Scholar
B. R. Whiteet al.,
“Bedside optical imaging of occipital resting-state functional connectivity in neonates,”
NeuroImage, 59
(3), 2529
–2538
(2012). http://dx.doi.org/10.1016/j.neuroimage.2011.08.094 NEIMEF 1053-8119 Google Scholar
M. Imaiet al.,
“Functional connectivity of the cortex of term and preterm infants and infants with Down’s syndrome,”
NeuroImage, 85
(1), 272
–278
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.04.080 NEIMEF 1053-8119 Google Scholar
H. Atsumoriet al.,
“Development of wearable optical topography system for mapping the prefrontal cortex activation,”
Rev. Sci. Instrum., 80
(4), 043704
(2009). http://dx.doi.org/10.1063/1.3115207 RSINAK 0034-6748 Google Scholar
T. Funaneet al.,
“Synchronous activity of two people’s prefrontal cortices during a cooperative task measured by simultaneous near-infrared spectroscopy,”
J. Biomed. Opt., 16
(7), 077011
(2011). http://dx.doi.org/10.1117/1.3602853 JBOPFO 1083-3668 Google Scholar
X. Cuiet al.,
“A quantitative comparison of NIRS and fMRI across multiple cognitive tasks,”
NeuroImage, 54
(4), 2808
–2821
(2011). http://dx.doi.org/10.1016/j.neuroimage.2010.10.069 NEIMEF 1053-8119 Google Scholar
S. Sasaiet al.,
“A NIRS-fMRI study of resting state network,”
NeuroImage, 63
(1), 179
–193
(2012). http://dx.doi.org/10.1016/j.neuroimage.2012.06.011 NEIMEF 1053-8119 Google Scholar
S. Heinzelet al.,
“Variability of (functional) hemodynamics as measured with simultaneous fNIRS and fMRI during intertemporal choice,”
NeuroImage, 71
(1), 125
–134
(2013). http://dx.doi.org/10.1016/j.neuroimage.2012.12.074 NEIMEF 1053-8119 Google Scholar
D. A. Boaset al.,
“Diffuse optical imaging of brain activation: approaches to optimizing image sensitivity, resolution, and accuracy,”
NeuroImage, 23
(1), S275
–288
(2004). http://dx.doi.org/10.1016/j.neuroimage.2004.07.011 NEIMEF 1053-8119 Google Scholar
I. Tachtsidiset al.,
“Measurement of frontal lobe functional activation and related systemic effects: a near-infrared spectroscopy investigation,”
Adv. Exp. Med. Biol., 614 397
–403
(2008). http://dx.doi.org/10.1007/978-0-387-74911-2_44 AEMBAP 0065-2598 Google Scholar
I. Tachtsidiset al.,
“Investigation of frontal cortex, motor cortex and systemic haemodynamic changes during anagram solving,”
Adv. Exp. Med. Biol., 614 21
–28
(2008). http://dx.doi.org/10.1007/978-0-387-74911-2_3 AEMBAP 0065-2598 Google Scholar
I. Tachtsidiset al.,
“False positives in functional near-infrared topography,”
Adv. Exp. Med. Biol., 645 307
–314
(2009). http://dx.doi.org/10.1007/978-0-387-85998-9_46 AEMBAP 0065-2598 Google Scholar
T. Takahashiet al.,
“Influence of skin blood flow on near-infrared spectroscopy signals measured on the forehead during a verbal fluency task,”
NeuroImage, 57
(3), 991
–1002
(2011). http://dx.doi.org/10.1016/j.neuroimage.2011.05.012 NEIMEF 1053-8119 Google Scholar
T. J. Germonet al.,
“Sensitivity of near infrared spectroscopy to cerebral and extra-cerebral oxygenation changes is determined by emitter-detector separation,”
J. Clin. Monit. Comput., 14
(5), 353
–360
(1998). http://dx.doi.org/10.1023/A:1009957032554 1387-1307 Google Scholar
L. Minatiet al.,
“Intra- and extra-cranial effects of transient blood pressure changes on brain near-infrared spectroscopy (NIRS) measurements,”
J. Neurosci. Methods, 197
(2), 283
–288
(2011). http://dx.doi.org/10.1016/j.jneumeth.2011.02.029 JNMEDT 0165-0270 Google Scholar
F. Alettiet al.,
“Deep and surface hemodynamic signal from functional time resolved transcranial near infrared spectroscopy compared to skin flowmotion,”
Comput. Biol. Med., 42
(3), 282
–289
(2012). http://dx.doi.org/10.1016/j.compbiomed.2011.06.001 CBMDAW 0010-4825 Google Scholar
S. Kohriet al.,
“Quantitative evaluation of the relative contribution ratio of cerebral tissue to near-infrared signals in the adult human head: a preliminary study,”
Physiol. Meas., 23
(2), 301
–312
(2002). http://dx.doi.org/10.1088/0967-3334/23/2/306 PMEAE3 0967-3334 Google Scholar
P. Smielewskiet al.,
“Clinical evaluation of near-infrared spectroscopy for testing cerebrovascular reactivity in patients with carotid artery disease,”
Stroke, 28
(2), 331
–338
(1997). http://dx.doi.org/10.1161/01.STR.28.2.331 SJCCA7 0039-2499 Google Scholar
Q. Zhanget al.,
“Adaptive filtering for global interference cancellation and real-time recovery of evoked brain activity: a Monte Carlo simulation study,”
J. Biomed. Opt., 12
(4), 044014
(2007). http://dx.doi.org/10.1117/1.2754714 JBOPFO 1083-3668 Google Scholar
S. Kohnoet al.,
“Removal of the skin blood flow artifact in functional near-infrared spectroscopic imaging data through independent component analysis,”
J. Biomed. Opt., 12
(6), 062111
(2007). http://dx.doi.org/10.1117/1.2814249 JBOPFO 1083-3668 Google Scholar
E. Kirilinaet al.,
“The physiological origin of task-evoked systemic artefacts in functional near infrared spectroscopy,”
NeuroImage, 61
(1), 70
–81
(2012). http://dx.doi.org/10.1016/j.neuroimage.2012.02.074 NEIMEF 1053-8119 Google Scholar
T. Funaneet al.,
“Quantitative evaluation of deep and shallow tissue layers’ contribution to fNIRS signal using multi-distance optodes and independent component analysis,”
NeuroImage, 85
(1), 150
–165
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.02.026 NEIMEF 1053-8119 Google Scholar
J. F. Cardoso,
“Blind signal separation: statistical principles,”
Proc. IEEE, 86
(10), 2009
–2025
(1998). http://dx.doi.org/10.1109/5.720250 IEEPAD 0018-9219 Google Scholar
T. Katuraet al.,
“Extracting task-related activation components from optical topography measurement using independent components analysis,”
J. Biomed. Opt., 13
(5), 054008
(2008). http://dx.doi.org/10.1117/1.2981829 JBOPFO 1083-3668 Google Scholar
H. Watanabeet al.,
“General to specific development of functional activation in the cerebral cortexes of 2- to 3-month-old infants,”
NeuroImage, 50
(4), 1536
–1544
(2010). http://dx.doi.org/10.1016/j.neuroimage.2010.01.068 NEIMEF 1053-8119 Google Scholar
G. Tagaet al.,
“Spatiotemporal properties of cortical haemodynamic response to auditory stimuli in sleeping infants revealed by multi-channel near-infrared spectroscopy,”
Philos. Trans. A Math. Phys. Eng. Sci., 369
(1955), 4495
–4511
(2011). http://dx.doi.org/10.1098/rsta.2011.0238 1364-503X Google Scholar
Y. Fukuiet al.,
“Monte Carlo prediction of near-infrared light propagation in realistic adult and neonatal head models,”
Appl. Opt., 42
(16), 2881
–2887
(2003). http://dx.doi.org/10.1364/AO.42.002881 APOPAI 0003-6935 Google Scholar
G. Tagaet al.,
“Effects of source-detector distance of near infrared spectroscopy on the measurement of the cortical hemodynamic response in infants,”
NeuroImage, 38
(3), 452
–460
(2007). http://dx.doi.org/10.1016/j.neuroimage.2007.07.050 NEIMEF 1053-8119 Google Scholar
P. van der Zeeet al.,
“Experimentally measured optical pathlengths for the adult head, calf and forearm and the head of the newborn infant as a function of inter optode spacing,”
Adv. Exp. Med. Biol., 316 143
–153
(1992). http://dx.doi.org/10.1007/978-1-4615-3404-4_17 AEMBAP 0065-2598 Google Scholar
M. S. Beauchampet al.,
“The developmental trajectory of brain-scalp distance from birth through childhood: implications for functional neuroimaging,”
PLoS One, 6
(9), e24981
(2011). http://dx.doi.org/10.1371/journal.pone.0024981 1932-6203 Google Scholar
H. H. Jasper,
“The ten twenty electrode system of the International Federation,”
Electroencephalogr. Clin. Neurophysiol., 10 371
–375
(1958). ECNEAZ 0013-4694 Google Scholar
M. Matsuiet al.,
“Referential framework for transcranial anatomical correspondence for fNIRS based on manually traced sulci and gyri of an infant brain,”
Neurosci. Res., 80 55
–68
(2014). http://dx.doi.org/10.1016/j.neures.2014.01.003 0168-0102 Google Scholar
F. Homaeet al.,
“The right hemisphere of sleeping infant perceives sentential prosody,”
Neurosci. Res., 54
(4), 276
–280
(2006). http://dx.doi.org/10.1016/j.neures.2005.12.006 0168-0102 Google Scholar
G. TagaK. Asakawa,
“Selectivity and localization of cortical response to auditory and visual stimulation in awake infants aged 2 to 4 months,”
NeuroImage, 36
(4), 1246
–1252
(2007). http://dx.doi.org/10.1016/j.neuroimage.2007.04.037 NEIMEF 1053-8119 Google Scholar
D. T. Delpyet al.,
“Estimation of optical pathlength through tissue from direct time of flight measurement,”
Phys. Med. Biol., 33
(12), 1433
–1442
(1988). http://dx.doi.org/10.1088/0031-9155/33/12/008 PHMBA7 0031-9155 Google Scholar
L. Gagnonet al.,
“Further improvement in reducing superficial contamination in NIRS using double short separation measurements,”
NeuroImage, 85
(1), 127
–135
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.01.073 NEIMEF 1053-8119 Google Scholar
R. B. Saageret al.,
“Two-detector corrected near infrared spectroscopy (C-NIRS) detects hemodynamic activation responses more robustly than single-detector NIRS,”
NeuroImage, 55
(4), 1679
–1685
(2011). http://dx.doi.org/10.1016/j.neuroimage.2011.01.043 NEIMEF 1053-8119 Google Scholar
T. Funaneet al.,
“Dynamic phantom with two stage-driven absorbers for mimicking hemoglobin changes in superficial and deep tissues,”
J. Biomed. Opt., 17
(4), 047001
(2012). http://dx.doi.org/10.1117/1.JBO.17.4.047001 JBOPFO 1083-3668 Google Scholar
E. OkadaD. T. Delpy,
“Near-infrared light propagation in an adult head model. I. Modeling of low-level scattering in the cerebrospinal fluid layer,”
Appl. Opt., 42
(16), 2906
–2914
(2003). http://dx.doi.org/10.1364/AO.42.002906 APOPAI 0003-6935 Google Scholar
G. E. Strangmanet al.,
“Scalp and skull influence on near infrared photon propagation in the Colin27 brain template,”
NeuroImage, 85
(1), 136
–149
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.04.090 NEIMEF 1053-8119 Google Scholar
T. Yamadaet al.,
“Separation of fNIRS signals into functional and systemic components based on differences in hemodynamic modalities,”
Plos One, 7
(11), e50271
(2012). http://dx.doi.org/10.1371/journal.pone.0050271 1932-6203 Google Scholar
R. A. Fisher,
“Frequency distribution of the values of the correlation coefficient in samples from an indefinitely large population,”
Biometrika, 10
(4), 507
–521
(1915). BIOKAX 0006-3444 Google Scholar
R. A. Fisher,
“On the ‘Probable Error’ of a coefficient of correlation deduced from a small sample,”
Metron, 1 3
–32
(1921). 0026-1424 Google Scholar
M. A. Franceschiniet al.,
“Hemodynamic evoked response of the sensorimotor cortex measured noninvasively with near-infrared optical imaging,”
Psychophysiology, 40
(4), 548
–560
(2003). http://dx.doi.org/10.1111/psyp.2003.40.issue-4 PSPHAF 0048-5772 Google Scholar
BiographyTsukasa Funane is a researcher at Central Research Laboratory, Hitachi, Ltd., Japan. Since 2006, he has been a member of the research group working on optical topography, optical brain function monitoring technology based on near-infrared spectroscopy. His main responsibilities include basic research on new measurement methods as well as hardware developments and its application to human brain study, such as social cognitions. Fumitaka Homae is an associate professor in the Department of Language Sciences, Tokyo Metropolitan University, Japan. His research focuses on developmental brain science of language. His research interests also include structural and functional development of global cortical networks in infancy. Hama Watanabe is a project associate professor at the Graduate School of Education, the University of Tokyo, Japan. She is currently involved in studies for infant behavioral and neurological developments. She contributes over broad topics ranging behavioral science, psychological science, cognitive science, developmental science, and neuroscience. Masashi Kiguchi has studied various optical measurements: nonlinear spectroscopy, time-resolved spectroscopy, near-field spectroscopy, and near-infrared spectroscopy, and his background is physics and laser spectroscopy. He has studied the problems related to the principle of near-infrared spectroscopy (NIRS) measurement and has been taking the lead in the development of new techniques for observing brain activities to open new research fields and in basic studies for putting them to practical use. Gentaro Taga is a professor at the Graduate School of Education, the University of Tokyo, Japan. His research broadly focuses on the early development of the brain and behavior in humans. He is a pioneer in dynamical system modeling of the neuro-musculo-skeletal system for human locomotion and applying functional NIRS imaging for infant studies. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||