|
|
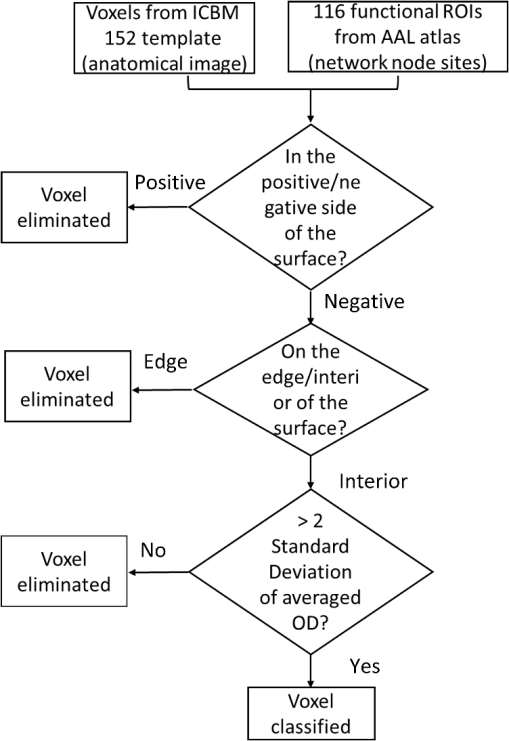
1.IntroductionAtlas-guided diffuse optical tomography (atlas-DOT) has undergone rapid development in recent years. Custo et al.1 initially introduced the concept of atlas-DOT, after which it was further developed with high-density optodes and successfully validated using computational simulations and functional magnetic resonance imaging (fMRI) performed on human subjects.2–4 Atlas-DOT utilized a finite element technique with either subject-specific models3,4 or a standard magnetic resonance imaging (MRI) brain template (such as ICBM 256) in the forward calculation.5,6 A significant improvement of accuracy in image reconstruction and source localization has been achieved with atlas-DOT.7 More recently, atlas-DOT has been utilized together with the general linear model analysis, which further extended its feasibility of measuring hemodynamic changes in complex brain tasks. Several groups have reported consistent and accurate results of brain hemodynamic changes under visual stimulation,7 speech,8 and risk decision-making.5 It is well known that the human brain is naturally organized into groups of networks, with anatomical brain regions involved in either individualized processing or integration with other brain regions, to accomplish different functions. The brain networks experience changes in their properties under a variety of conditions, such as taking on different physical or mental tasks, facing strong emotions,9 or undergoing normal aging.10–12 Thus, noninvasive mapping of the human brain’s structural and functional connectivity enables researchers to better understand the architecture of the brain and to clearly reveal connectivity changes in entire brain networks. Increasingly, many neuroimaging techniques, such as fMRI, diffusion MRI, electroencephalographic (EEG), and DOT, have been recently used to investigate large-scale brain networks.13–17 Accordingly, researchers have developed corresponding mathematical frameworks that can better model and analyze brain networks for a variety of imaging modalities. Graph theory analysis (GTA) is one of the analytical methods developed to examine large-scale complex brain networks.13–15,18,19 It can provide an uncomplicated and yet powerful mathematical means to characterize the brain networks’ topological properties. Specifically, GTA is a mathematical method for the analysis of complex network systems; it finds applications in the modeling of interactions in information, social, and biological systems. Recently, GTA has been reported in the neuroimaging literature to analyze resting-state functional connectivity (RSFC).18,20,21 It depicts a brain network as a graph with different anatomical brain regions portrayed as nodes and interactions between brain regions represented as links. Based on GTA, a number of studies have found alterations in network parameters in aged patients. Liu et al.11 conducted a resting-state positron emission tomography (PET) study in young and aged patients and reported a decrease in network efficiency for older patients, who also maintained a higher clustering coefficient than younger subjects. This result indicates that the brain network becomes more random in nature as humans grow older. The reduction in gray matter as a result of aging has also been associated with a decrease in global efficiency and increase in path length for older subjects.22,23 The activities of the default mode network have been also reported to decrease in older adults,24 indicating changes in the network’s properties along with aging. Furthermore, the aging process has been implicated in alterations in modularity and number of hubs of the brain network.25,26 In the field of near-infrared spectroscopy (NIRS), a few recent studies have combined GTA with channel-wise NIRS and revealed topological organization and architecture of large-scale, resting-state human brain cortical networks.27–29 However, no report combines atlas-DOT with GTA in brain network research. In addition, little validation exists for the atlas-DOT technique in the assessment of RSFC. The difficulties in using atlas-DOT to form brain networks result from the difficulties of graph formation, the initial necessary step in GTA. Instead of using a channel-wise approach, a data set of resting-state atlas-DOT images is four-dimensional (4-D), consisting of multiple three-dimensional (3-D) cortical volumes in a time sequence. Because a reconstructed atlas-DOT image often consists of a large number of voxels, it is computationally expensive and lacks an appropriate means to generate the connectivity matrices needed in GTA. Routinely used techniques to extract regions of interest (ROIs) in fMRI analysis (such as threshold-, anatomy-, and function-based ROI30) cannot be directly applied to atlas-DOT. In this paper, we introduced an automated voxel classification (AVC) approach to facilitate graph formation for atlas-DOT images by identifying or grouping unregulated voxel distribution. By using a subject-averaged brain template (ICBM 152),31 we demonstrated that our method could fit for any brain size. We then provided a solution of voxel classification based on the automated anatomical labeling (AAL) of activations in 116 segmented structures (AAL 116).32 In this way, the graph formation was guided by the anatomical structure of the brain. To test our approach, we conducted a test–retest assessment by measuring RSFC in 15 young adults. Moreover, we quantified age-related changes in the chosen brain network and compared our results with previously published PET-derived reports. Both reproducibility and cross-modality comparison indicated that our AVC with atlas-DOT is an efficient and feasible tool for assessing functional brain networks. 2.Methods and MaterialsWe developed a three-section data processing strategy to perform atlas-DOT with AVC. As represented in Fig. 1, the red, yellow, and blue dashed lines enclose the three major steps: data acquisition and preprocessing, atlas-DOT, and graph formation with AVC, respectively. After generating the brain network matrices (or adjacency matrices), we performed the group-level GTAs and statistical analyses to test the reproducibility of brain connectivity in the young and older adults. All of our analyses were processed using MATLAB® 2015b (The MathWorks, Inc., Natick, Massachusetts). Fig. 1The work flow for AVC used with atlas-DOT. The first step (marked by the red dashed line box) includes the ICBM 152 brain template generation, data acquisition, and data preprocessing. The second step (marked by the yellow dashed line box), also referred to as atlas-DOT, includes finite element computation for brain atlas, forward modeling, and image reconstruction. The third step (marked by the blue dashed line box) illustrates graph formation with AVC, which includes volume segmentation using predefined AAL with 116 brain regions (AAL 116), voxel classification and ROIs generation, and formation of an adjacency matrix by cross-correlation. To validate our approach, a routine GTA with statistic analysis was followed up to access the local and global network features. Refer to details in the text. 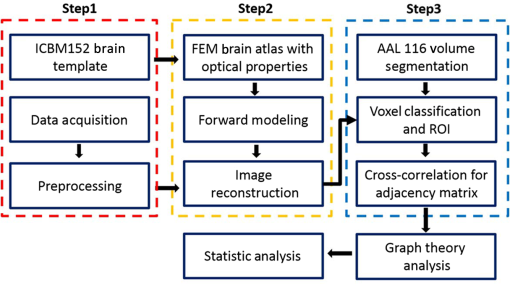 2.1.Participants, Protocols, Data Acquisition, and Preprocessing2.1.1.Participants and protocolsThe study participants comprised 15 young adults (ages 25 to 43) recruited from the University of Texas at Arlington and 21 older adults (ages 65 to 92) recruited from the city of Arlington, Texas. All subjects were right-handed with normal visual ability. No subjects reported any known diseases such as musculoskeletal, neurological, visual, or cardiorespiratory dysfunctions. Written consent forms were obtained for all participants before the protocol started. The Institutional Review Board of the University of Texas at Arlington approved this study. Before the measurement, all participants were instructed to sit comfortably in a quiet room with eyes closed and with minimal body movements. Our protocol included two repeated measurements on the young adults with a time separation of two weeks. We also included one-time recordings for 21 older adults; however, four older adults’ data were excluded from data analysis because of head movements, requested recesses, or failure to complete the protocols. All the measurements of 10-min RSFC were performed during daytime in the same experiment room, maintained at a constant temperature, humidity, and light intensity. 2.1.2.Data acquisitionDuring the protocol, a continuous wave, multichannel functional NIRS (fNIRS) brain imaging system (Cephalogics, Massachusetts) was applied to each subject’s forehead to record cortical hemodynamic alterations during resting state [see Fig. 2(a)]. The wavelengths used to calculate changes of oxygenated hemoglobin () and deoxygenated hemoglobin () were 750 and 850 nm. The sensor array consisted of 18 pairs of two light sources each and 18 detectors with the nearest interoptode distance of 2.5 cm and the second-nearest interoptode distance of 3.5 cm, forming 75 measurement channels and covering the forehead entirely [see Fig. 2(b)]. To find anatomical locations of the optodes, the international 10–20 system of electrodes placement was used to coregister the locations of sources and detectors to a standardized brain atlas. In addition, the reference points of nasion (Na), inion (In), top (Cz), left ear (A1), and right ear (A2) in the 10–20 system were measured using a 3-D digitizer to perform the brain size normalization in subsequent steps. The region of detection covered approximately surface area in the forehead, which mainly covered the Brodmann areas 9, 10, and 11. The detected fNIRS signals were acquired at a sampling frequency of 10.8 Hz. Fig. 2Experimental setup and optode/probe geometry: (a) optical probe placement on a partcipant’s forehead and (b) optical optode geometry. Open circles represent source locations and solid circles represent detector locations. The source–detector (SD) separation is 2.5 cm, which indicates that the nearest SD distance is 2.5 cm and the second-nearest SD is 3.5 cm. The total number of SD pairs is 75. 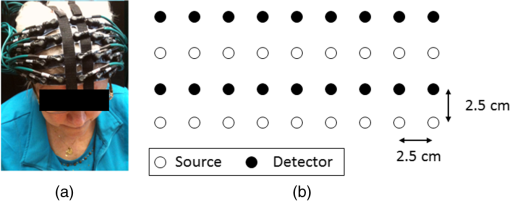 2.1.3.Data preprocessingFor data preprocessing, bandpass filtering and a global autocorrelation process were sequentially applied to the channel-wise raw fNIRS data. Specifically, the bandpass filter chosen in this study was 0.02 to 0.3 Hz, as suggested by a previous study, to eliminate the physiological noise generated by heartbeat and respirations.5 A global signal referencing approach was applied to the channel-wise fNIRS signals to further eliminate the unmodeled global physiology or structure noise.33 Specifically, for each participant, averaging over 75-channel time sequences generated a global signal. Then, we applied an autocorrelation approach between the globally averaged signal and signals from each individual channel to perform global regression. Any channel-wise signal strongly correlated with the global signal was considered to contain strong artifacts. Global referencing was performed in these channels by deducting the global signal from the individual channel signals.33 2.2.Brain Imaging with Atlas-Guided Diffuse Optical TomographyOur previous study5 and many others from the literature2,4,6 extensively introduced the atlas-DOT algorithm. We briefly describe the method as follows. First, the predesigned ICBM152 brain template including 176,192 voxels was generated using the MRI structure scan, an average brain image taken from 152 subjects.31 The size of the brain template was . A careful segmentation was performed to separate the scalp, skull, gray, and white matters. A finite element surface mesh was generated for each segment and later grown inside each of the four segments (i.e., scalp, skull, gray, and white matters) to form a complete volume, constituted with tetrahedrons. A total of 179,162 vertexes were near-uniformly distributed in Montreal Neurological Institute (MNI) space. Considering the differences of optical properties in the real human brain, each segment was assigned a specific set of optical properties suggested by a previous study.2 Table 1 provides the optical properties of different brain layers. Table 1Optical properties of head tissues for modeling.
Second, spatial coregistration and head size normalization were performed in order to minimize the confounding by the brain size variation generated as a result of normal aging. As indicated in Sec. 2.1.2, we performed location reference measurements before acquiring data for both young and older adults. Specifically, we performed an affine transformation34 between the standard 10–20 system landmarks in ICBM152 MNI space and the 3-D digitizer measurements of subject-specific locations.5,31 The coordinates of projected optodes for each individual were then normalized in the standard MNI space in the ICBM 152 brain atlas. Figure 3(a) shows the normalized optical source and detector locations plotted with the ICBM 152 cortical image template. The red spots represent the locations of light sources and the blue spots represent the locations of light detectors. Fig. 3Illustration of the AVC algorithm. (a) Locations of optical optodes (sources: red dots; detectors: blue dots) projected on the ICBM 152 brain cotrical template. (b) Spatial distribution of DOT measurement sensitivity after FEM forward modeling, based on the normalized optical optode geometries. (c) Averaged locations of AAL 116 regions by colored spots. They have been separated into six networks: default (yellow), frontal–parietal (light blue), occipital (blue), sensorimotor (navy), cingulo-opercular or limic (orange), and cerebellum (dark red). (d) A 3-D view of the AAL 116 ROIs and the surfaces based on Ref. 32. (e) AVC categorizes all atlas-DOT voxels interrogated by the fNIRS probes in this study into six different brain regions as represented by different colors. (f) The locations or coordinates of the 34 ROIs within the atlas-DOT measured in this study. (g) Time-sequence plots of 34 nodal values from one subject (subject 01) during a 10-min resting-state period. The -axis represents time, and the -axis represents the oxygenated hemoglobin changes of 34 nodes. 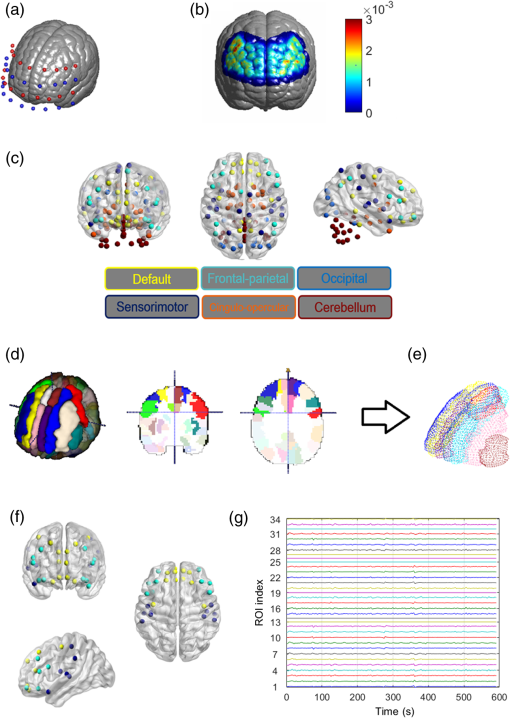 Third, the forward modeling of light propagation in the ICBM152 brain template was performed based on the Rytov approximation.35,36 The light sensitivity matrix (or Jacobin matrix) was created by performing the finite element estimation of light propagation inside the ICBM152 brain atlas using a finite element method (FEM)-based MATLAB® package called NIRFAST.37 The optical probe geometry shown in Fig. 3(a) generated 75 measurement channels. A depth compensation algorithm with a tested compensation factor of 1.7 was further applied to the sensitivity matrix to project the reconstructed DOT images at more accurate brain layers.38 By visually checking the light sensitivity in the forehead [Fig. 3(b)], we observed that the measurements primarily covered Brodmann 9, 10, 11, and 46, including the rostrolateral prefrontal cortex, dorsolateral prefrontal cortex, and parts of the parietal regions. Last, to recover spatial–temporal changes of light absorptions resulting from brain resting-state hemodynamic activities, we implemented a 3-D image reconstruction approach repeatedly at different time points. We performed inverse image reconstruction using the Moore–Penrose generalized approach with Tikhonov regularization.39 The reconstructed images were 4-D matrices with 176,192 spatial voxels and 6480 temporal points (10 min with a sampling rate of 10.8 Hz). Consequently, determining absorption changes at two wavelengths led to reconstructed images of relative changes in ΔHbO concentrations, based on spectral decomposition of the extinction coefficients for both wavelengths.40 2.3.Graph Formation with Automated Voxel ClassificationTo perform GTA, we first needed to generate a proper functional connectivity matrix called graph formation, in which the ROIs are classified as nodes, and interactions between nodes are termed edges.18 Edges in brain networks could be either binary or weighted. Binary edges indicate the presence or absence of interactions (“0” for absence or “1” for presence), while weighted edges represent interaction strength. Using a threshold can remove weak edges. Edges in functional brain studies are usually quantified by calculating the correlation coefficient between two nodes. In functional brain network analysis, graph formation is usually performed directly from the measured data. Niu et al.29 introduced a method using the channel-wise time sequence data as the graph formation input. However, this method does not work for atlas-DOT, because time-dependent data in atlas-DOT are not given in channels but in voxels. In graph formation for fMRI measurements, voxel-wise time-sequence data are often classified or grouped into certain clusters based on either anatomical locations or predefined functional ROIs.41 The method we applied here shares a similar idea. Following a widely accepted anatomical template, namely, the AAL atlas,32 we presented or projected the 116 anatomical ROIs on the ICBM 152 brain atlas (see more details in Sec. 2.3.1). We then performed AVC to identify respective anatomical clusters on the ICBM 152-based atlas-DOT. Next, we calculated Pearson correlation coefficients by performing cross-correlations of the time series between each pair of identified clusters to create the adjacency matrices. Figure 4 shows a flowchart describing the overall selection process of AVC. 2.3.1.Anatomical automated labeling with volume segmentationBecause the ICBM 152 brain atlas provided only an average anatomical image but lacked detailed anatomical features,32 we had to use the AAL 116-based ROIs to facilitate or define node definitions in our study (as shown in Fig. 4). While the AAL 116 ROIs were originally derived from a single-subject T1-weighted MRI, those ROIs could be still projected or normalized on the ICBM 152 brain template.32 Relatively, there was little difference in 3-D views of corresponding AAL ROIs on the ICBM 152 brain template and the single-subject T1-weighted brain template. More explanation is given in Appendix A. Specifically, we separated the human brain into 116 anatomical ROIs based on the landmarks of cortical and subcortical structures, as marked by the AAL 116 ROIs. Further, these 116 regions were categorized into six brain networks based on Ref. 22: default mode (yellow), sensorimotor (navy blue), frontal–parietal lobe (light blue), occipital lobe (blue), cingulo-opercular/limbic lobe (orange), and cerebellum (dark red), as identified and presented in Fig. 3(c) using the respective colors. BrainNet Viewer software42 was used to view or image the three planes. 2.3.2.Voxel classification to identify the region of interestTo perform graph formation within voxel-wise atlas-DOT images, we developed an AVC algorithm to assign each DOT cortical voxel to an appropriate ROI, namely one of the AAL 116 neuroanatomical regions. The selected ROIs then served as graph nodes for graph formation, followed by cross-correlations of different time sequences between each pair of ROI-based nodes. First, both the ICBM152 and the AAL116 templates were transformed to the same coordinate space. Here, we chose the MNI coordinates as our standard space coordinate reference. As indicated in Sec. 2.2, an affine transformation was performed on the ICBM 152 atlas after the FEM forward calculation, resulting in standard MNI coordinates for each voxel (, , , , where is the total number of voxels). The MNI coordinates of the original AAL 116 template were obtained from a previous study.32 Second, the surfaces of AAL 116 template were extracted using the software ITK,43 shown in Fig. 3(d), and exported them as vertices (e.g., , , , , where is the total number of voxels) and faces (triangles). To efficiently and correctly identify or classify whether the brain voxels were within a certain AAL region, we followed the algorithm called “point-in-polyhedron problem.”44 Specifically, the first operation, the orientation operation, tested whether a point falls to either positive or negative sides of a triangle within surface. The second operation, point classification operation, classified whether a point is on one of the triangle’s edges or in its interior. After the voxel classification, we were able to group or classify all the voxels in an atlas-DOT brain image into 116 anatomical regions. For example, Fig. 3(e) shows all atlas-DOT voxels on the cortical template classified into six different brain regions, represented by different colors. Second, the voxels were further selected based on the forward model of light diffusion theory. Because the optical measurements in our study did not cover the whole brain volume, the sensitivity of light interrogation in brain regions deeper than 2 to 3 cm rapidly decayed due to strong light scattering with increasing depth. Therefore, we applied a threshold to eliminate all the voxels whose optical sensitivity (quantified as optical density, OD) was less than two standard deviations of the total OD within an anatomical region. After this selection, the remaining voxels were combined with the results of voxel classification from the previous step. Using this two-step operation, we were able to quantify or classify 34 ROIs within the 116 anatomical regions. For graph formation, these 34 ROIs were regarded as nodes. Table 2 lists network names, anatomical regions or descriptions, labels, and brain atlas coordinates of these 34 nodes. In addition, Fig. 3(f) plots the averaged coordinates within the 34 ROIs. These 34 nodes in our study were included partially within the default mode network (yellow), frontal–parietal network (light blue), and sensorimotor network (navy blue). Each time sequence, averaged over all the voxels within an ROI (i.e., each of the 34 nodes), was treated as the input of node information to create the adjacency matrix. As an example, Fig. 3(g) shows a time-sequence plot of 34 nodal ΔHbO values from one subject (subject 01). Specifically, the -axis represents time within 10 min of a resting-state period and the -axis represents oxygenated hemoglobin changes with the node index from 1 to 34. Table 2Node positions based on AAL atlas used in this study
2.3.3.Cross-correlation to generate adjacency matrixFormer studies using fNIRS-based GTA to investigate brain networks suggested that correlations of brain hemodynamic changes or fluctuations among different brain regions could represent brain functional connectivity.27–29 The same strategy or rationale was applied in this study to nodal time series, using the 10-min resting-state fNIRS measurements to establish the adjacency matrix for each participant. The cross-correlation between each pair of nodes was performed for the given time series, as shown in Fig. 3(g), and Pearson correlation coefficients () were computed to form a adjacency correlation matrix [see Fig. 5(a)]. Note that the color in Fig. 5(a) denotes the values of the correlation coefficients: blue indicates a low-correlation coefficient and red represents a high-correlation coefficient. The adjacency matrix was further converted into a binarized matrix by setting a threshold. The correlation coefficient between node and was set to 1, if the correlation value was larger than the given threshold and 0 otherwise [see Fig. 5(b)]. The edge was defined as functional connections ( to represent a significant correlation between two nodes or = 0 to represent no significant correlation). Fig. 5(a) The adjacency matrix generated by cross-correlation of values at 34 nodes from one human subject. The - and -axes represent the node numbers; red color represents a value with high correlation between the temporal profiles of two nodes, while blue color represents a low-correlation coefficient. (b) A binary matrix thresholded by a middle sparsity of 0.25. Note that the diagonal elements were set to be zero since they were not the actual two-channel correlations. (c) A 3-D view of spatial representation of nodes and edges generated to show the RSFC. The connected gray lines represent the fNIRS-derived RSFC measured in this study, whereas the unconnected dots show the ROIs classified by our AVC algorithm. 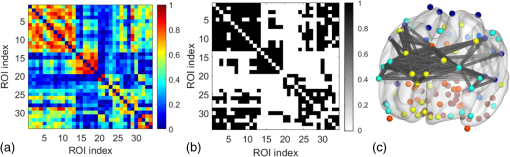 In principle, different settings or selections on the cross-correlation threshold will result in a different binarized matrix. In this study, we applied different thresholds to the adjacency matrix to obtain a sequence of binary matrices. Specifically, the sparsity-based approach was utilized as suggested by Niu et al.29 Sparsity () for a fixed graph was defined as the number of current existing edges in this graph divided by the maximum possible number of edges in the current graph. In this study, the range from 0.1 to 0.5 (i.e., , ) was chosen to be the standard threshold sequence, as a previous study reported.11 Then, the threshold sequence was applied to the adjacency matrix to generate 41 binarized network matrices for each subject. Figure 5(c) shows an example from one subject’s data, revealing a spatial representation of nodes and edges from one binary matrix with a middle threshold (). Colors of nodes represent different brain anatomical regions, whereas gray lines between two nodes represent respective connections. Actually, the connected lines represent the fNIRS-derived RSFC measured in this study, whereas the unconnected dots show the ROIs that our AVC algorithm classified. 2.4.Generating Brain Network Matrices Using Graph Theory AnalysisBased on the adjacency matrices, we further quantified the resting-state brain network parameters using GTA. Graph theory metrics for functional brain networks were calculated under global and local network characteristics. In this study, we did not include all of the global and local parameters given by GTA because of the processing load required and the purpose of this study. We focused on demonstrating our AVC by using a few major parameters affected by age effects in a previous study.11 Specifically, a set of global parameters investigated were global network metrics, including (i) clustering coefficient (), (ii) shortest path length (), (iii) global efficiency (), (iv) normalized clustering coefficient (), (v) normalized characteristic path length (), and (vi) small-world (). For the local parameters, we focused on the hub information, including (i) nodal degree, (ii) nodal efficiency, and (iii) betweenness centrality. Appendix B includes detailed descriptions of these graph metrics. 2.5.Statistical AnalysisWe confirmed our classification algorithm by performing two statistical analyses. First, we performed a test–retest analysis to evaluate the reproducibility of quantification of the brain networks between two measurement visits in 15 young adults. We also performed a second analysis to compare the differences of global and local features between young and older adults. Specifically, we computed global efficiency (), shortest path length (), clustering coefficient (), normalized clustering coefficient (), normalized characteristic path length (), and small-world coefficient () for all the groups. Normality was performed on all measurements, and data without normality were converted to -values () transformation before further analysis. 2.5.1.Evaluating the reproducibility of current approachThe evaluation of reproducibility was performed in both brain connectivity and graph theory metrics in 15 young adults. First, group-level functional connectivity adjacency matrices from the two visits were compared. Specifically, Pearson correlation coefficients were computed for all pixels in the RSFC matrices. Then a linear fitting between two visits were performed for all the pixels. Second, global (i.e., , , , , , and ) and local [i.e., nodal efficiency (), nodal degree (), and betweenness centrality ()] parameters of the two visits were compared. Since the fNIRS measurements were sequentially performed, a two-sample -test was taken at each sparsity threshold for all global parameters, resulting in a range of significant tests along sparsity. A criterion of was selected to define statistical significance in this study. 2.5.2.Analysis of the age effect on the global and local parametersWe further evaluated our method by performing an age-related analysis between 15 young adults and 21 older adults in both global (, , , , , and ) and local (nodal efficiency, nodal degree, and betweenness centrality) brain network characteristics. The two-sample -tests for the global metrics were performed between two groups along all sparsity values (). In addition, regions of hubs were compared in nodal efficiency (), nodal degree (), and betweenness centrality (). 3.ResultsWe performed careful analysis to test the reproducibility and feasibility of our AVC method. The reproducibility analysis of the method was based on the measurements from young adults, with each of the individuals having participated in the fNIRS recordings twice within a period of two weeks. The method’s feasibility analysis tested the age effect on the brain networks by comparing them between the young and older adults. 3.1.Resting-State Functional Connectivity MatricesTo check the reproducibility of our approach, 10-min rest-state fNIRS measurements were taken from young adults () twice in two separate experiments. A total number of 75 channels allowed us to cover and interrogate most of the prefrontal regions [see Figs. 3(a) and 3(b)]. Our newly developed AVC algorithm was used resulting in 34 nodes within the cortical regions reconstructed using atlas-DOT [see Figs. 3(f) and 5(c)]. All the 10-min temporal profiles from the 34 nodes were extracted [see Fig. 3(g)] and used to create RSFC adjacency matrices, whose elements or pixels were filled with respective Pearson correlation coefficients () between each pair of the 34 nodes. Figures 6(a) and 6(b) show such RSFC matrices group-averaged over the 15 young adults for visits 1 and 2, respectively. Visual inspection showed that the two RSFC matrices were consistent with each other in overall distribution of correlation patterns and correlation strengths. More quantitatively, we compared these two matrices by plotting the two separate sets against each other, as shown in Fig. 6(c). The comparison demonstrated a linear correlation between the two visits . These results demonstrated that our AVC algorithm was strongly reproducible when used with atlas-DOT to obtain RSFC matrices. Fig. 6RSFC matrices averaged over young adults at (a) visit 1 and (b) visit 2. Each data pixel marks the Pearson correlation coefficient () calculated between each pair of 34 nodes. Note that the diagonal elements were set to be zero since they were not actual two-channel correlations. (c) A linear relationship is illustrated between two data sets of young adults at visit 1 (-axis) and visit 2 (-axis). The red line is the fitted line of the two groups , indicating a significant correlation between the two visits. 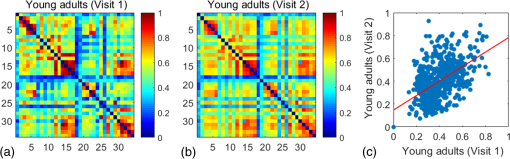 It is known that GTA allows for the analysis and categorization of topological measures of brain networks into two groups: (a) global network metrics: small-world properties (e.g., clustering coefficient , characteristic path length , normalized clustering coefficient , normalized characteristic path length , and small-world ), efficiency parameters (local efficiency and global efficiency ), modularity , hierarchy , and assortativity and (b) nodal characteristics: nodal degree, nodal efficiency, and nodal betweenness.27 We followed the same grouping strategy to present our analysis in Sec. 3.2. 3.2.Global Network Characteristics and Small-World FeaturesTo further evaluate the reproducibility of our algorithm, we quantified and compared the global network and small-world features of the brain networks between the two visits of young adults and between young and older adults. Figures 7(a) to 7(c) show the respective values of clustering coefficient (), shortest path length (), and global efficiency () of young adults obtained from two separate measurements along with the corresponding readings from the older adults. Two-sample -tests for each of the global brain network parameters were performed at each sparsity value from 0.1 to 0.5, as a previous study suggested.11 Fig. 7Global characteristics of the GTA. (a)–(c) indicate the clustering coefficient (), shortest path length (), and global efficiency () quantified from young adults in visit 1 (solid circles) and visit 2 (open circles) and from older adults (pluses). The dashed line on the bottom of panel (a) and (c) marks the sparsity ranges where significant differences in respective network parameters exist between two visits (or measurements) of young adults. The solid lines on the bottom of each panel represent the sparsity ranges where significant differences in respective network parameters exist between young and older adults. (d)–(f) indicate the small-world characteristics, including the normalized characteristic path lengths (), normalized clustering coefficient (), and small-worldness () quantified from young adults in visit 1 (solid circles) and visit 2 (open circles) and from older adults (pluses), respectively. The solid lines on the bottom of (d) to (f) have the same meaning as those in (a) to (c). 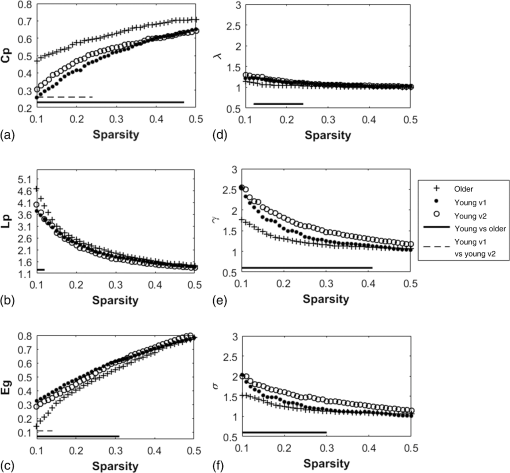 3.2.1.Significant differences in global features between two visits of young adultsFigures 7(a) to 7(c) present the values of clustering coefficient (), shortest path length (), and global efficiency () determined from two visits of young adults. Overall, results illustrated a high reproducibility of the global characteristics of the brain networks. As Figs. 7(a) and 7(c) show, significant differences in and between visits 1 and 2 existed only within a short range of the sparsity. showed no significant difference within the given sparsity range between visits 1 and 2. 3.2.2.Significant differences in global features between young and older adultsFor better comparison, we also plotted global features of the brain networks from older adults together with those from the young adults. Figures 7(a) and 7(c) clearly show age-related changes in and . Quantitatively, the corresponding global features of young adults from the first visit were used to carry out statistical assessment. Specifically, older adults revealed a significantly larger clustering coefficient () within the sparsity range between 0.1 and 0.47 [see Fig. 7(a)]. On the other hand, young adults had a significantly larger global efficiency () than older adults within the sparsity range of 0.1 to 0.31 [see Fig. 7(c)]. Meanwhile, we observed no significant difference in shortest path length () between the two age groups [see Fig. 7(b)]. Our findings were consistent with a previous study11 that investigated resting-state brain networks in 115 young adults () and 110 older adults () using PET. Their results indicated a global efficiency decline with an increase in clustering coefficient in older adults.11 3.2.3.Small-world functional networkFunctional networks of the human brain have small-world characteristics.21,45 Compared to random networks, the small-world characteristics in the human brain include large local connectivity and an approximately identical shortest path length between any two nodes in the network. Figures 7(d) to 7(f) show the normalized characteristic path length (), normalized clustering coefficient (), and small-worldness () taken from young adults in visit 1, visit 2, and from older adults, respectively. We observed that in all three groups approached 1 with sparsity to 0.5 [see Fig. 7(d)], and so did . Noticeably, no significant difference existed in , , and between two visits of young adults, revealing excellent reproducibility in the small-world features between the two measurements. Regarding age effects on the small-world characteristics of the brain networks, our results, furthermore, demonstrated that the young adults have significantly larger than older adults over a range of sparsity (). In addition, the young adults have clearly exhibited larger values of () and () over a large range of sparsity than older adults. 3.3.Reproducibility in Local Graph ParameterIn this study, network hubs were also quantified by nodal degree (), nodal efficiency (), and betweenness centrality (). The nodal metrics were constructed at a sparsity threshold of 0.16, as a previous study suggested,11 to ensure that the networks of both young and older adult groups had the same number of nodes and edges. The hubs were then selected for each of the three nodal parameters (i.e., , , and ) with respective values larger than one standard deviation of the corresponding average values over all nodes, as previous studies suggested.27,29 Figures 8(a) to 8(c) demonstrate axial or top views of the hubs determined from the young adults in visits 1 and 2 and from the older adults for all three nodal metrics of degree (), efficiency (), and betweenness centrality (). As Niu et al.29 indicated, the betweenness centrality was considered as the major reference for the hub measurements.9 Fig. 8Hubs determined from (a) young adults in visit 1, (b) young adults in visit 2, and (c) older adults. The hubs are presented based on nodal degree (), nodal efficiency (), and betweenness centrality (). Yellow dots represent the hubs within the default mode network, light blue dots represent the hubs within the frontal–parietal network, and dark blue dots represent those within the sensorimotor network. [See Figs. 3(f) and 5(c).] 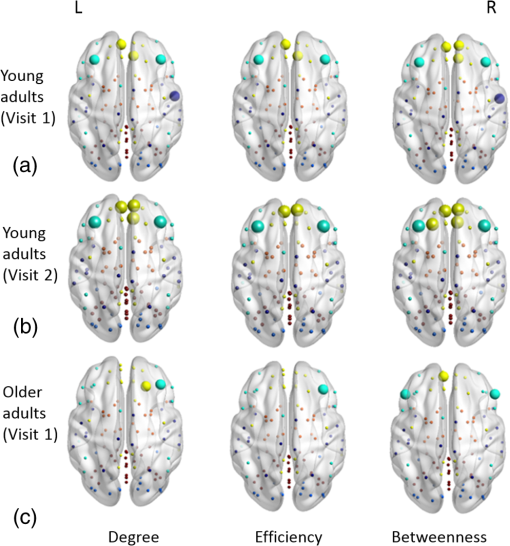 3.3.1.Hubs determined within young adultsAs Figs. 8(a) and 8(b) show, the young adults presented a few consistent and bilateral hub positions in the prefrontal cortex region obtained in both of the two visits. Specifically, we observed bilateral hub locations in middle frontal gyrus (MFG.L, MFG.R), superior frontal gyrus (SFGmed.L, SFGmed.R), and right anterior cingulate gyrus (ACG.R) for the betweenness centrality in both visits. Young adult visit 1 (not visit 2) exhibited the hub position in the right rolandic operculum (ROL.R), and young adult visit 2 exhibited the hub position in the left superior frontal gyrus (SFGdor.L). The nodal degree and nodal efficiency indicated fewer hubs than the betweenness centrality. The current study used the betweenness centrality as the marker of the hubs. 3.3.2.Hubs found differently between young and older adultsIn the older adult group, as shown in Fig. 8(c), two, one, and three hubs were identified under nodal degree, nodal efficiency, and betweenness centrality, respectively. In contrast, five, four, and six hubs were identified from young adults in visit 1; similarly, five, four, and six hubs were also found in visit 2. In particular, the two visits identified very similar hubs (four in nodal degree, four in nodal efficiency, and five in betweenness centrality). These results revealed an overall decline of hub numbers in the older adult group. Furthermore, changes in regional nodal characteristics between the two age groups were also noted. Specifically, hubs in default mode networks were observed only on the right side of the middle frontal gyrus (MFG.R) and the left dorsolateral region of the medial superior frontal gyrus (SFGmed.L) in the older adult group, whereas both sides of the middle frontal gyrus (MFG.L and MFG.R) and the dorsolateral region (SFGmed.L and SFGmed.R) were observed in the young adult group. In addition, the bilateral hub pairs (MFG.L, MFG.R, SFGmed.L, and SFGmed.R) were observed in young adults, while unilateral hub (SFGmed.L) was observed in older adults for betweenness centrality. Table 3 summarizes and presents all the hub locations. Table 3Regions of hubs in two age groups.
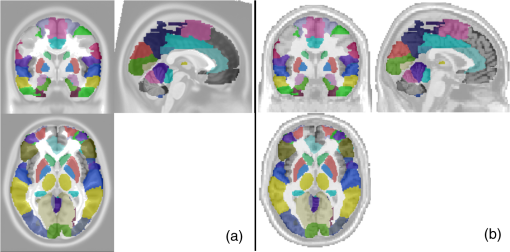
*Marks hub locations from one visit of young adults.**Marks hub locations from two visits of young adults.○Marks hub locations from one visit of older adults. 4.Discussion4.1.Development of Automated Voxel Classification With Atlas-Guided Diffuse Optical TomographyAtlas-DOT is an imaging method recently established within the NIRS-based neuroimaging field.1–8,46 GTA is a network analysis tool that has been commonly used in functional neuroimaging to study brain connectivity and networks.13–15,18,19 To date, however, atlas-DOT and GTA have not been used jointly to extract or derive large-scale brain connectivity and networks from the NIRS measurements because of computational challenges posed by atlas-DOT. Specifically, graph formation in GTA is a difficult and computationally expensive process for atlas-DOT because of the large amount of voxels in atlas-DOT images. The major novelty in our current work was the development of AVC with atlas-DOT, which affords image processing speed and localization accuracy in optical brain imaging. Such an automated classification is critical because it helps to extract and identify cerebral hemodynamic signals from appropriately matched anatomical locations, thus improving the accuracy of interpretations of local and global brain activities. Commonly used methods for registering NIRS data on a human brain template are often based on the EEG 10–20 system,47–49 which is not directly applicable to the 3-D atlas-DOT images. In addition, routinely used voxel classification in fMRI is based on an affine transformation, which converts the voxel space to MNI space, followed by a voxel matching with a standard MNI structure template.41 However, this approach works only when the voxels in the brain images are homogeneously distributed or have the same equal voxel size. This approach is not applicable to atlas-DOT for several reasons. First, because atlas-DOT is FEM based, different input parameters (such as the head size, geometry, and shape) to the atlas model would result in a different total number and distribution of voxels in the data space because of the nature of FEM computation and methodology. Second, the spatial distribution of optical densities used for atlas-DOT is variable and heterogeneous, and it depends on the optical properties of the human head with multiple layers. Such heterogeneous multilayer structures result in an unpredictable number and pattern of voxels in a certain region. Therefore, the spatial classification method used by fMRI cannot be transferred directly for voxel classification in atlas-DOT. Furthermore, no universal template can be generated to match the FEM voxels versus specific anatomical structures because each FEM voxel model must be determined according to the optical probe geometries or setups. Thus, we were motivated to address this need and develop the current technique, which is a comprehensive yet efficient solution for the atlas-DOT voxel classification. The principle of AVC was based on the “point-in-polyhedron problem”44 approach. The basic idea was to identify whether a specific voxel was (i) inside, (ii) outside, or (iii) just on a particular surface. Specifically, our approach focused on the location of each atlas-DOT voxel with respect to the surface of several predefined brain structures. Thus, no matter what the voxel distribution was, once the voxel coordinates in the MNI space were assigned, the classification of DOT voxels to an appropriate anatomical ROI would be automatically achieved. Because this morphological operation or classification with atlas-DOT is efficient, it can be applied to other atlas-DOT-based brain imaging applications. 4.2.Test–Retest Reliability of Resting-State Functional Connectivity Using Automated Voxel Classification With Atlas-Guided Diffuse Optical TomographyAnother innovative aspect of this study was that it demonstrated the good reproducibility of AVC with atlas-DOT in assessing RSFC. We found strong consistency between two matrices of RSFC taken from two separate visits in two weeks (Fig. 6). A recent report by Niu et al.27 revealed a better reproducibility of the RSFC correlation, with a higher test–retest correlation coefficient than ours . One possible reason for this difference resulted from the fact that our two test–retest measurements were separated by two weeks, whereas the measurements in Ref. 27 were obtained from only the same set of measurements taken at two time periods. Furthermore, the global network matrices obtained from the two visits have shown an overall high consistency [see Figs. 7 and 8], as two-sample -tests also proved. Specifically, except for the clustering coefficient in the sparsity range from , global network features demonstrated good reproducibility along the sparsity range of 0.1 to 0.5. This finding confirmed that during a resting state, the prefrontal cortex activities were noticeably repeatable.27 Similar hub locations observed in both visits also validated the reproducibility of local network matrices. For example, we observed bilateral network hubs in middle frontal and superior frontal gyrus within the three nodal metrics of degree, efficiency, and betweenness centrality in both visits. All these brain regions were consistent with those reported and observed previously by fMRI and PET brain network studies.11,18,21 4.3.Comparison of Age Effect on Resting-State Functional Connectivity Using Automated Voxel Classification With Atlas-Guided Diffuse Optical TomographyAnother innovative aspect of this study was that it clearly revealed the age effect on the resting-state brain network observed by combining AVC with atlas-DOT. Although both fMRI and PET have observed and demonstrated age effects on the brain networks by Lieu et al.11 and Meunier et al.,12 no report on this topic exists using optical methods. In this study, we further investigated partial default mode networks and frontal–parietal networks of young adults (ages 25 to 43) and older adults (ages 65 to 92). Our findings were compatible with previous studies,11,50,51 which showed a higher global efficiency in young adults than in older adults. It is believed that the anatomical alterations of aging are responsible for these changes, especially after the age of 65.52 Our present results also revealed a difference in clustering coefficient between the older and young adults. Similar findings from previous reports11,25 suggest that a possible reorganization of frontal network occurs as people age. It is also suggested that information processes are less economical in older adults, especially in the frontal and temporal cortical and subcortical regions.25 Several other reports have also indicated an increase of shortest path length with an increase in age resulting from the loss of long-distance connections and interconnected hubs.25,53 However, we found no differences in this network metric, possibly because our optical probe setup covered a limited brain region (i.e., prefrontal), leading to a shortage of long-distance brain network measurements. Finally, we have observed a significant decrease in small-worldness in older adults, implying a degeneration process, in good agreement with previous studies.11,12,18 Our findings have also identified several hubs of the brain networks located within the frontal cortical regions in both age groups, as reported by several previous publications.11,18 Meanwhile, a decrease in the number of hubs in the older adults was observed as compared to the young adults. Specifically, the nodal betweenness in default mode regions (such as middle frontal gyrus and superior frontal gyrus) was diminished in the older adults. 4.4.Consideration of Spatial LeakageIn recent years, much research in the context of EEG- and MEG-based RSFC has focused on possible confounding effects resulting from spatial leakage.54–56 This concern is essential when the measurement data are reconstructed into source-space. Since such connectivity images or maps are formed by solving an ill-posed inverse problem, there are inherent correlations among adjacent vertices. Each reconstructed brain vertex can be expressed as a linear combination of the channel-space data. This confounding issue is more severe for correction in MEG- and EEG-based connectivity studies than fNIRS-derived brain network mappings because the optical sensitivity falls off much faster (within 1 to 2 cm) than MEG ( to 6 cm).55 It is still worthwhile to estimate the spatial leakage that may cause misleading conclusions in our study. For this purpose, we have added Appendix C that describes our estimation approach to quantify the spatial leakage. Appendix C presents the results and draws a basic conclusion. This extra section mainly implies that the spatial leakage is short-distanced, and the nearby region within 1 to 2 cm could be affected. Future investigation is needed in order to understand better how this distance-related artifactual connectivity can largely affect the brain network interpretations. One possible solution for correction of spatial leakage is to obtain a measurement of spatial leakage artifact by an “empty-room” recording and then to normalize the actual human brain data with respect to the artifactual data.55 Another approach, named geometric correction scheme, was based on subtraction of the spatial leakage geometry model from the reconstructed image data and approved efficient to minimize the spatial leakage effect.54 4.5.Limitations of this Study and Suggestions for Future WorksBecause the light penetration depth through the human brain tissue (not including scalp nor skull) is not more than 1 cm, the atlas-DOT technique can image only the brain regions mainly in gray matter.57 Thus, our atlas-DOT-based graph formation can provide brain network nodes and other metrics within only 1 cm below the cortical surface. Imaging a deeper brain layer requires larger separations between optical sources and detectors, which may lead to a significant signal-to-noise ratio decrease.58 Suggestions for future studies include simultaneous or parallel measurements of EEG recordings and fNIRS, which may help us better understand the relationship between neuroelectrical and cerebrovascular functions and their networks. Another limitation of this study was associated with physiological noises mixed within the measured fNIRS signals. Such noises were caused by superficial layers of the human head, such as the scalp and skull.59,60 To reduce these noises, we applied a global referencing method, which seemed promising because our results showed high consistency and reproducibility. However, we suggest further examining and confirming the noise effect using other approaches. Future studies could use a computational method such as independent component analysis27,61 or an experimental approach such as short-distance measurement regression2,4,7,8,46,57 to minimize global signal or noise effects. It is also highly suggested to increase the number of optodes in future studies in order to cover the entire cortical region. Because this study focused primarily on the development of AVC, we chose to cover only the frontal regions of the subjects’ heads to achieve efficient data acquisition. The cortical region detected in this study was only partial: about one-quarter of the entire brain cortical area. Node extraction in this study contained 34 nodal regions (out of 116 regions) in the AAL 116 atlas. Thus, further studies intending to rigorously determine resting-state functional brain networks using atlas-DOT should increase the number of recording optodes and cover the entire cortical region. In this way, researchers can optimally utilize AVC with atlas-DOT and obtain the most accurate results of resting-state brain network analysis. 5.ConclusionIn this study, we introduced an AVC approach that can facilitate graph formation for atlas-DOT images by grouping unregulated or unequally sized FEM voxels into anatomically meaningful ROIs within the human brain. To have an accurate human brain template for use with AVC, we chose a subject-averaged brain template (i.e., ICBM 152)31 along with the AAL 116 template32 to mark the brain ROIs. Thus, the graph formation was guided by the anatomical structure of the brain. To demonstrate the usefulness of AVC, we conducted a test–retest assessment by measuring RSFC in 15 young adults and quantified age-related changes in the measured brain network. Our results derived from AVC with atlas-DOT were in good agreement with those reported by a previous PET study. Overall, this study confirmed that AVC with atlas-DOT has great potential to serve as an efficient and feasible tool to quantify voxel-wise, resting-state functional brain networks. AppendicesAppendix A:Is There Any ROI Difference Shown on the ICBM 152 Atlas and on the Single-Subject T1-Weighted AAL Brain Template?Originally, the AAL 116 brain template was derived from a single subject whose brain was scanned 27 times by T1-weighted MRI. Specifically, these MRI images were coregistered and averaged to yield high detailed brain landmarks in MNI space (namely, Single_subj_T1) (see Ref. 32). On the other hand, the ICBM 152 brain was created by acquiring MRI over 152 human brains, followed by unbiased nonlinear averages of the scan brain images (see Ref. 31). To compare ROIs images based on these two brain templates, we have plotted AAL-based 116 ROIs in Fig. 9 using (a) the ICBM 152 brain template [see Fig. 9(a)] and (b) the single-subject T1-weighted brain template, which was used to segment AAL 116 ROIs [Fig. 9(b)]. Note that there was very little difference in ROI locations and/or anatomical variations, except that 3-D views of the ROIs on ICBM 152 are smoother because of better spatial resolution of the brain template. Appendix B:Brain Network Metrics18For a defined network or graph, , there are nodes and edges; the global and local network metrics are the output of the GTA. Using the GRETNA program, we computed the following graph metrics in our study:
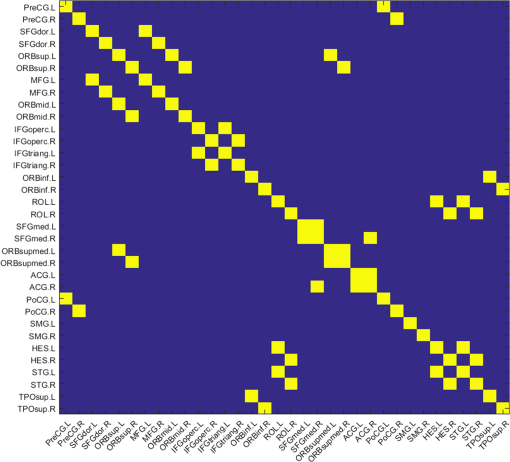
Appendix C:Estimation of Spatial Leakage in fNIRS-Derived Functional Connectivity MappingTo estimate the spatial leakage in fNIRS-derived functional connectivity, we performed a seed-based “spatial leakage” test on a set of empty-room data, following the method given in Refs. 54 and 55. A set of 6-min fNIRS temporal data were recorded while the optical probe was placed on a “dummy head” in a dark room. The same image reconstruction process was performed on the raw dummy data to obtain the voxel-wised time sequences. A seed region was chosen on the left side of the prefrontal lobe [as marked by black circle in Fig. 10(a)]. The Pearson’s correlation coefficients, , between the seed region to all other voxels of the brain were computed and rendered on the 3-D human brain template [see Fig. 10(a)], and these values versus corresponding Euclidean distances are also plotted in Fig. 10(b). This figure shows that the correlation coefficient drops rapidly within a distance of approximately 16 to 20 mm, revealing that most of voxels affected by the “spatial leakage” are within 1.5 to 2 cm from the seed site. Fig. 10(a) Pearson’s correlation coefficients, , between the seed region (marked by the black circle) and all other voxels covered by the optical optodes, were computed and rendered on the 3-D dummy brain template. (b) It plots the values versus corresponding Euclidean distances from the seed to other voxels. 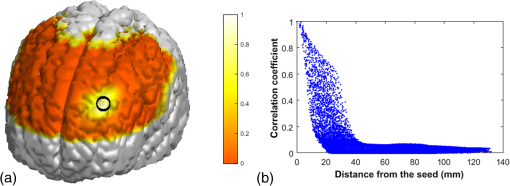 To specifically estimate the effect of “spatial leakage” in our current study, we investigated individual Euclidean distances across all the 34 AAL ROIs that we used in the network formation and presented the results in Fig. 11. In this figure, - and -axes are the identified 34 ROIs, while blue color represents the ROI-to-ROI distance being larger than 20 mm and yellow color indicates the distance shorter than 20 mm. We can see that 22 out of 561 () ROI-to-ROI distances are shorter than 20 mm, implying that the “spatial leakage” effect is not too much significant (about 3.9% of total ROIs). However, the “spatial leakage” effect could vary depending on locations of the seed because the light propagation in the human brain is not homogeneous. More rigorous analysis should be performed in future studies to confirm this preliminary conclusion. AcknowledgmentsThe authors acknowledge the support in part from the 2012 Southern Nursing Research Society Research Award. ReferencesA. Custo et al.,
“Anatomical atlas-guided diffuse optical tomography of brain activation,”
NeuroImage, 49 561
–567
(2010). http://dx.doi.org/10.1016/j.neuroimage.2009.07.033 NEIMEF 1053-8119 Google Scholar
A. T. Eggebrecht et al.,
“A quantitative spatial comparison of high-density diffuse optical tomography and fMRI cortical mapping,”
NeuroImage, 61 1120
–1128
(2012). http://dx.doi.org/10.1016/j.neuroimage.2012.01.124 NEIMEF 1053-8119 Google Scholar
Y. Zhan et al.,
“Image quality analysis of high-density diffuse optical tomography incorporating a subject-specific head model,”
Front. Neuroenerg., 4 6
(2012). http://dx.doi.org/10.3389/fnene.2012.00006 Google Scholar
R. J. Cooper et al.,
“Validating atlas-guided DOT: a comparison of diffuse optical tomography informed by atlas and subject-specific anatomies,”
NeuroImage, 62 1999
–2006
(2012). http://dx.doi.org/10.1016/j.neuroimage.2012.05.031 NEIMEF 1053-8119 Google Scholar
Z. J. Lin et al.,
“Atlas-guided volumetric diffuse optical tomography enhanced by generalized linear model analysis to image risk decision-making responses in young adults,”
Hum. Brain Mapp., 35 4249
–4266
(2014). http://dx.doi.org/10.1002/hbm.v35.8 HBRME7 1065-9471 Google Scholar
F. Tian and H. Liu,
“Depth-compensated diffuse optical tomography enhanced by general linear model analysis and an anatomical atlas of human head,”
NeuroImage, 85
(Pt. 1), 166
–180
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.07.016 NEIMEF 1053-8119 Google Scholar
X. Wu et al.,
“Quantitative evaluation of atlas-based high-density diffuse optical tomography for imaging of the human visual cortex,”
Biomed. Opt. Express, 5 3882
–3900
(2014). http://dx.doi.org/10.1364/BOE.5.003882 BOEICL 2156-7085 Google Scholar
M. S. Hassanpour et al.,
“Mapping cortical responses to speech using high-density diffuse optical tomography,”
NeuroImage, 117 319
–326
(2015). http://dx.doi.org/10.1016/j.neuroimage.2015.05.058 NEIMEF 1053-8119 Google Scholar
D. Yurgelun-Todd,
“Emotional and cognitive changes during adolescence,”
Curr. Opin. Neurobiol., 17 251
–257
(2007). http://dx.doi.org/10.1016/j.conb.2007.03.009 COPUEN 0959-4388 Google Scholar
G. Gong et al.,
“Age- and gender-related differences in the cortical anatomical network,”
J. Neurosci., 29 15684
–15693
(2009). http://dx.doi.org/10.1523/JNEUROSCI.2308-09.2009 JNRSDS 0270-6474 Google Scholar
Z. Liu et al.,
“Changes in topological organization of functional PET brain network with normal aging,”
PLoS One, 9 e88690
(2014). http://dx.doi.org/10.1371/journal.pone.0088690 POLNCL 1932-6203 Google Scholar
D. Meunier et al.,
“Age-related changes in modular organization of human brain functional networks,”
NeuroImage, 44 715
–723
(2009). http://dx.doi.org/10.1016/j.neuroimage.2008.09.062 NEIMEF 1053-8119 Google Scholar
U. Braun et al.,
“Test–retest reliability of resting-state connectivity network characteristics using fMRI and graph theoretical measures,”
NeuroImage, 59 1404
–1412
(2012). http://dx.doi.org/10.1016/j.neuroimage.2011.08.044 NEIMEF 1053-8119 Google Scholar
M. Rubinov and O. Sporns,
“Complex network measures of brain connectivity: uses and interpretations,”
NeuroImage, 52 1059
–1069
(2010). http://dx.doi.org/10.1016/j.neuroimage.2009.10.003 NEIMEF 1053-8119 Google Scholar
D. J. Kim et al.,
“Disturbed resting state EEG synchronization in bipolar disorder: a graph-theoretic analysis,”
Neuroimage Clin., 2 414
–423
(2013). http://dx.doi.org/10.1016/j.nicl.2013.03.007 Google Scholar
A. T. Eggebrecht et al.,
“Mapping distributed brain function and networks with diffuse optical tomography,”
Nat. Photonics, 8 448
–454
(2014). http://dx.doi.org/10.1038/nphoton.2014.107 NPAHBY 1749-4885 Google Scholar
Q. Cao et al.,
“Probabilistic diffusion tractography and graph theory analysis reveal abnormal white matter structural connectivity networks in drug-naive boys with attention deficit/hyperactivity disorder,”
J. Neurosci., 33 10676
–10687
(2013). http://dx.doi.org/10.1523/JNEUROSCI.4793-12.2013 JNRSDS 0270-6474 Google Scholar
E. Bullmore and O. Sporns,
“Complex brain networks: graph theoretical analysis of structural and functional systems,”
Nat. Rev. Neurosci., 10 186
–198
(2009). http://dx.doi.org/10.1038/nrn2575 NRNAAN 1471-003X Google Scholar
X. H. Liao et al.,
“Functional brain hubs and their test–retest reliability: a multiband resting-state functional MRI study,”
NeuroImage, 83 969
–982
(2013). http://dx.doi.org/10.1016/j.neuroimage.2013.07.058 NEIMEF 1053-8119 Google Scholar
C. J. Stam and J. C. Reijneveld,
“Graph theoretical analysis of complex networks in the brain,”
Nonlinear Biomed. Phys., 1 3
(2007). http://dx.doi.org/10.1186/1753-4631-1-3 Google Scholar
Y. He and A. Evans,
“Graph theoretical modeling of brain connectivity,”
Curr. Opin. Neurol., 23 341
–350
(2010). http://dx.doi.org/10.1097/WCO.0b013e32833aa567 Google Scholar
N. U. Dosenbach et al.,
“Prediction of individual brain maturity using fMRI,”
Science, 329 1358
–1361
(2010). http://dx.doi.org/10.1126/science.1194144 SCIEAS 0036-8075 Google Scholar
E. Bullmore and O. Sporns,
“The economy of brain network organization,”
Nat. Rev. Neurosci., 13 336
–349
(2012). http://dx.doi.org/10.1038/nrn3214 NRNAAN 1471-003X Google Scholar
J. S. Damoiseaux et al.,
“Reduced resting-state brain activity in the ‘default network’ in normal aging,”
Cereb. Cortex, 18 1856
–1864
(2008). http://dx.doi.org/10.1093/cercor/bhm207 Google Scholar
S. Achard and E. Bullmore,
“Efficiency and cost of economical brain functional networks,”
PLoS Comput. Biol., 3 e17
(2007). http://dx.doi.org/10.1371/journal.pcbi.0030017 Google Scholar
S. Achard et al.,
“A resilient, low-frequency, small-world human brain functional network with highly connected association cortical hubs,”
J. Neurosci., 26 63
–72
(2006). http://dx.doi.org/10.1523/JNEUROSCI.3874-05.2006 JNRSDS 0270-6474 Google Scholar
H. Niu et al.,
“Test–retest reliability of graph metrics in functional brain networks: a resting-state fNIRS study,”
PLoS One, 8 e72425
(2013). http://dx.doi.org/10.1371/journal.pone.0072425 POLNCL 1932-6203 Google Scholar
H. Niu and Y. He,
“Resting-state functional brain connectivity: lessons from functional near-infrared spectroscopy,”
Neuroscientist, 20 173
–188
(2014). http://dx.doi.org/10.1177/1073858413502707 NROSFJ 1073-8584 Google Scholar
H. Niu et al.,
“Revealing topological organization of human brain functional networks with resting-state functional near infrared spectroscopy,”
PLoS One, 7 e45771
(2012). http://dx.doi.org/10.1371/journal.pone.0045771 POLNCL 1932-6203 Google Scholar
R. A. Poldrack,
“Region of interest analysis for fMRI,”
Soc. Cogn. Affect Neurosci., 2 67
–70
(2007). http://dx.doi.org/10.1093/scan/nsm006 Google Scholar
S. Cutini, P. Scatturin and M. Zorzi,
“A new method based on ICBM152 head surface for probe placement in multichannel fNIRS,”
NeuroImage, 54 919
–927
(2011). https://doi.org/http://www.bic.mni.mcgill.ca/ServicesAtlases/ICBM152NLin2009 NEIMEF 1053-8119 Google Scholar
N. Tzourio-Mazoyer et al.,
“Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain,”
NeuroImage, 15 273
–289
(2002). http://dx.doi.org/10.1006/nimg.2001.0978 NEIMEF 1053-8119 Google Scholar
G. K. Aguirre, E. Zarahn and M. D’Esposito,
“Empirical analyses of BOLD fMRI statistics. II. Spatially smoothed data collected under null-hypothesis and experimental conditions,”
NeuroImage, 5 199
–212
(1997). http://dx.doi.org/10.1006/nimg.1997.0264 NEIMEF 1053-8119 Google Scholar
A. K. Singh et al.,
“Spatial registration of multichannel multi-subject fNIRS data to MNI space without MRI,”
NeuroImage, 27 842
–851
(2005). http://dx.doi.org/10.1016/j.neuroimage.2005.05.019 NEIMEF 1053-8119 Google Scholar
T. Durduran et al.,
“Diffuse optics for tissue monitoring and tomography,”
Rep. Prog. Phys., 73 076701
(2010). http://dx.doi.org/10.1088/0034-4885/73/7/076701 RPPHAG 0034-4885 Google Scholar
S. R. Arridge and J. C. Schotland,
“Optical tomography: forward and inverse problems,”
Inverse Prob., 25 70
(2009). http://dx.doi.org/10.1088/0266-5611/25/12/123010 INPEEY 0266-5611 Google Scholar
H. Dehghani et al.,
“Near infrared optical tomography using NIRFAST: algorithm for numerical model and image reconstruction,”
Commun. Numer. Methods Eng., 25 711
–732
(2008). http://dx.doi.org/10.1002/cnm.v25:6 CANMER 0748-8025 Google Scholar
H. Niu et al.,
“Development of a compensation algorithm for accurate depth localization in diffuse optical tomography,”
Opt. Lett., 35 429
–431
(2010). http://dx.doi.org/10.1364/OL.35.000429 OPLEDP 0146-9592 Google Scholar
J. A. Fill and D. E. Fishkind,
“The Moore–Penrose generalized inverse for sums of matrices,”
SIAM J. Matrix Anal. Appl., 21 629
–635
(2000). http://dx.doi.org/10.1137/S0895479897329692 SJMAEL 0895-4798 Google Scholar
D. A. Boas and A. M. Dale,
“Simulation study of magnetic resonance imaging-guided cortically constrained diffuse optical tomography of human brain function,”
Appl. Opt., 44 1957
–1968
(2005). http://dx.doi.org/10.1364/AO.44.001957 APOPAI 0003-6935 Google Scholar
J. A. Etzel, V. Gazzola and C. Keysers,
“An introduction to anatomical ROI-based fMRI classification analysis,”
Brain Res., 1282 114
–125
(2009). http://dx.doi.org/10.1016/j.brainres.2009.05.090 BRREAP 0006-8993 Google Scholar
M. Xia, J. Wang and Y. He,
“BrainNet Viewer: a network visualization tool for human brain connectomics,”
PLoS One, 8 e68910
(2013). http://dx.doi.org/10.1371/journal.pone.0068910 POLNCL 1932-6203 Google Scholar
“National library of medicine insight segmentation and registration toolkit (ITK),”
(2016) https://itk.org May ). 2016). Google Scholar
J. Liu et al.,
“A new point containment test algorithm based on preprocessing and determining triangles,”
Comput. Aided Des., 42 1143
–1150
(2010). http://dx.doi.org/10.1016/j.cad.2010.08.002 CAIDA5 0010-4485 Google Scholar
D. S. Bassett and E. Bullmore,
“Small-world brain networks,”
Neuroscientist, 12 512
–523
(2006). http://dx.doi.org/10.1177/1073858406293182 NROSFJ 1073-8584 Google Scholar
M. S. Hassanpour et al.,
“Statistical analysis of high density diffuse optical tomography,”
NeuroImage, 85
(Pt. 1), 104
–116
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.05.105 Google Scholar
F. Tian et al.,
“Prefrontal responses to digit span memory phases in patients with post-traumatic stress disorder (PTSD): a functional near infrared spectroscopy study,”
Neuroimage Clin., 4 808
–819
(2014). http://dx.doi.org/10.1016/j.nicl.2014.05.005 Google Scholar
F. Tian et al.,
“Test–retest assessment of cortical activation induced by repetitive transcranial magnetic stimulation with brain atlas-guided optical topography,”
J. Biomed. Opt., 17 116020
(2012). http://dx.doi.org/10.1117/1.JBO.17.11.116020 JBOPFO 1083-3668 Google Scholar
D. Tsuzuki and I. Dan,
“Spatial registration for functional near-infrared spectroscopy: from channel position on the scalp to cortical location in individual and group analyses,”
NeuroImage, 85
(Pt. 1), 92
–103
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.07.025 NEIMEF 1053-8119 Google Scholar
C. L. Grady et al.,
“Age-related changes in brain activity across the adult lifespan,”
J. Cognit. Neurosci., 18 227
–241
(2006). http://dx.doi.org/10.1162/jocn.2006.18.2.227 JCONEO 0898-929X Google Scholar
T. Hedden et al.,
“Disruption of functional connectivity in clinically normal older adults harboring amyloid burden,”
J. Neurosci., 29 12686
–12694
(2009). http://dx.doi.org/10.1523/JNEUROSCI.3189-09.2009 JNRSDS 0270-6474 Google Scholar
R. Peters,
“Ageing and the brain,”
Postgrad. Med. J., 82 84
–88
(2006). http://dx.doi.org/10.1136/pgmj.2005.036665 Google Scholar
J. D. Power et al.,
“The development of human functional brain networks,”
Neuron, 67 735
–748
(2010). http://dx.doi.org/10.1016/j.neuron.2010.08.017 NERNET 0896-6273 Google Scholar
V. Wens et al.,
“A geometric correction scheme for spatial leakage effects in MEG/EEG seed-based functional connectivity mapping,”
Hum. Brain Mapp., 36 4604
–4621
(2015). http://dx.doi.org/10.1002/hbm.22943 HBRME7 1065-9471 Google Scholar
A. S. Ghuman, J. R. McDaniel and A. Martin,
“A wavelet-based method for measuring the oscillatory dynamics of resting-state functional connectivity in MEG,”
NeuroImage, 56 69
–77
(2011). http://dx.doi.org/10.1016/j.neuroimage.2011.01.046 NEIMEF 1053-8119 Google Scholar
B. T. Schmidt, A. S. Ghuman and T. J. Huppert,
“Whole brain functional connectivity using phase locking measures of resting state magnetoencephalography,”
Front. Neurosci., 8 141
(2014). http://dx.doi.org/10.3389/fnins.2014.00141 1662-453X Google Scholar
S. L. Ferradal et al.,
“Atlas-based head modeling and spatial normalization for high-density diffuse optical tomography: in vivo validation against fMRI,”
NeuroImage, 85
(Pt. 1), 117
–126
(2014). http://dx.doi.org/10.1016/j.neuroimage.2013.03.069 NEIMEF 1053-8119 Google Scholar
G. E. Strangman, Z. Li and Q. Zhang,
“Depth sensitivity and source–detector separations for near infrared spectroscopy based on the Colin27 brain template,”
PLoS One, 8 e66319
(2013). http://dx.doi.org/10.1371/journal.pone.0066319 POLNCL 1932-6203 Google Scholar
J. R. Goodwin, C. R. Gaudet and A. J. Berger,
“Short-channel functional near-infrared spectroscopy regressions improve when source–detector separation is reduced,”
Neurophotonics, 1 015002
(2014). http://dx.doi.org/10.1117/1.NPh.1.1.015002 Google Scholar
R. J. Cooper et al.,
“The utility of near-infrared spectroscopy in the regression of low-frequency physiological noise from functional magnetic resonance imaging data,”
NeuroImage, 59 3128
–3138
(2012). http://dx.doi.org/10.1016/j.neuroimage.2011.11.028 NEIMEF 1053-8119 Google Scholar
H. Zhang et al.,
“Test–retest assessment of independent component analysis-derived resting-state functional connectivity based on functional near-infrared spectroscopy,”
NeuroImage, 55 607
–615
(2011). http://dx.doi.org/10.1016/j.neuroimage.2010.12.007 NEIMEF 1053-8119 Google Scholar
BiographyLin Li received his BS degree in optoelectronics from the Huazhong University of Science and Technology in 2009 and his PhD in biomedical engineering from the University of Texas at Arlington, in 2015. Currently, he is a postdoctoral fellow at the University of California, Los Angeles. His current research interests include resting-state functional brain imaging, and multimodality neuronal disorders studies and algorithm development. Mary Cazzell received her BS and PhD in nursing from Marquette University and the University of Texas at Arlington, respectively. She is the director of Nursing Research and Evidence-Based Practice, Cook Children’s Medical Center, Fort Worth, Texas. Her research area focused on behavioral measurement of risk decision-making and identification of related neural correlates, using optical imaging, across the lifespan. Olajide Babawale received his bachelor of engineering degree in electrical and electronics engineering from Covenant University, Nigeria, and his master’s degree in biomedical engineering from the University of Texas at Arlington. He is a second-year doctoral student at the University of Texas at Arlington. His current research work includes multimodal neuroimaging using electroencephalographic (EEG) and functional near-infrared spectroscopy (fNIRS) modalities, EEG functional connectivity analysis using source-space imaging approaches, and fNIRS analysis of cognitively impaired children. Hanli Liu received her MS and PhD degrees in physics from Wake Forest University, followed by postdoctoral training in tissue optics at the University of Pennsylvania. She is a full professor of bioengineering at the University of Texas at Arlington. Her current expertise lies in the field of near-infrared spectroscopy of tissues, optical sensing for cancer detection, and diffuse optical tomography for functional brain imaging, all of which are related to clinical applications. |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||